Nov. 11, 2014 Press Release Biology Chemistry
For enterics, adaptability could be an Achilles heel
In research published in Nature Chemical Biology, scientists from RIKEN in Japan have discovered a surprisingly simple mechanism through which enterics can adjust to the very different oxygen environments inside the human gut and outside. This research, which was led by Shigeyuki Yokoyama and Wataru Nishii of the Structural Biology Laboratory, opens a new potential target against these bacteria, which are the most-frequently encountered causative microorganisms of infectious diseases. The family includes well-known symbionts and facultative or obligate pathogens such as E. coli, Klebsiella, Salmonella, Shigella and Yersinia pestis.
The team found that a subtle change in an enzyme called Lon, which is involved in the process of proteolysis, allows these bacteria to quickly adapt between the changing oxygen environments inside the gut—where there is practically no oxygen—and outside, where there is plenty. Proteolysis is an important process through which cells degrade unneeded proteins, and it must be tightly controlled to avoid cellular damage. The team discovered a small and surprisingly simple conformational change, based on a single disulfide bond, that acts as the switch, allowing the cell to go into a higher proteolysis mode when exposed to an oxygen-rich aerobic environment.
The team used three approaches. First, they analyzed the Lon protease using crystallography and showed that the enzyme can adopt both an oxidized and reduced form. They found that these states could switch in a reversible way, though the formation and reduction of a disulfide bond that caused the exit pore of the enzyme to either widen or narrow. They found that this small change had a dramatic impact on the enzyme's ability to carry out proteolysis.
Next, they investigated the enzymatic properties of Lon in solution, and found that its activity increased in high-oxygen conditions and decreased in conditions such as those in the gut. They identified the change in the exit pore size as the cause of the change.
Finally, they showed that the redox switch actually functions in living cells, demonstrating that cellular Lon activity was low in anaerobic and high in aerobic conditions, carefully regulating the double-edged sword of proteolysis, which can defend cells against external stress but harm them when unnecessary.
According to Wataru Nishii, the first author and one of the two corresponding authors of the studies, "We were very surprised by the fact that a tiny structural change, caused by the bonding and reduction of a single disulfide bond, had such a strong effect on molecular and cellular functions. This is the first example of an 'allosteric disulfide bond' that actually regulates the molecular function in normal physiological conditions. Though it remains a target for future investigation, it is an interesting possibility that such mechanisms may also exist in other proteins."
Shigeyuki Yokoyama, the other corresponding author, said, "Pathologically speaking, we have discovered a novel mechanism through which enterics can proliferate inside and outside the host's body. It was an unexpected, and could lead to a paradigm shift in our understanding of infectious diseases. It is tempting to imagine that the mechanism could become a new therapeutic target in infectious diseases, presumably by targeting the disulfide bond and/or the exit pore. Developing drugs would be quite important, since enterics cause a number of very familiar diseases that can be deadly, especially in weakened patients."
Reference
- Wataru Nishii, Mutsuko Kukimoto-Niino, Takaho Terada, Mikako Shirouzu, Tomonari Muramatsu, Masaki Kojima, Hiroshi Kihara, & Shigeyuki Yokoyama.
"A Redox switch shapes the Lon protease exit pore to facultatively regulate proteolysis" Nature Chemical Biology, In Press. 2014, doi: 10.1038/nchembio.1688
Contact
Distinguished Senior Scientist
Shigeyuki Yokoyama
Structural Biology Laboratory
Distinguished Senior Scientist Laboratories
Jens Wilkinson
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr@riken.jp
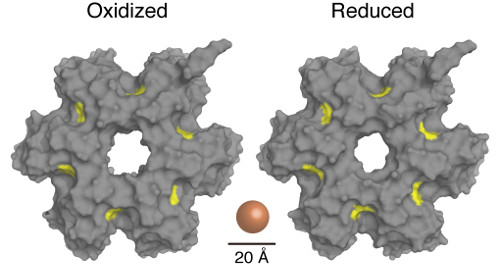
Image showing the structure of the enzyme in a reduced and oxidized environment
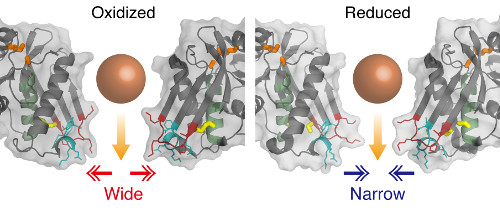
Image showing the structure near the exit pore
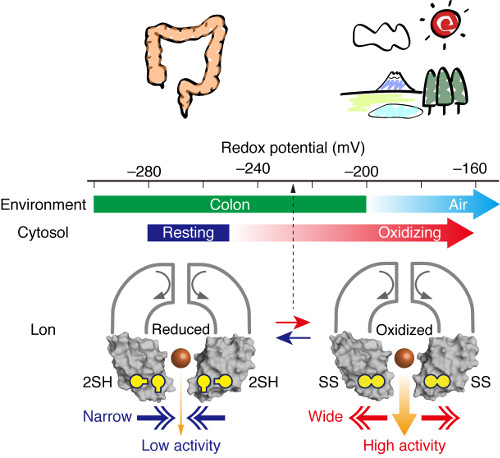
Schematic of how the enzyme changes in response to varying redox conditions
