Jan. 12, 2018 Press Release Biology
The brain's GPS has a buddy system
To be successful as a social animal, you need to know where you stand relative to others. Brain cells that perform precisely this function—locating the 'self' and others in space—have now been identified. In rats, the same brain area that stores the animal’s own location also maps the movements of other rats. Sometimes these representations are processed jointly by the same cells, depending on a rat’s goals and actions. This discovery, from Japan’s RIKEN Brain Science Institute, deepens our understanding of the hippocampus and its role as the brain’s positioning system.
It has been known for some time that the hippocampus maintains a mental map of space—in fact, the 2014 Nobel Prize in Physiology or Medicine was awarded precisely for this research. ‘Place cells’ and ‘grid cells’ in the hippocampus register the location of the brain’s owner in its environment, but until now, little was known about how the movements of others are tracked in the brain. Researchers put this to the test by observing the activity of hippocampal neurons in one rat (the ‘self’) watching another rat (the ‘other’) go through a simple T-maze. The self’s neurons registered what the other was doing and changed their responses based on the self’s location and subsequent actions. This study was published on January 11 in Science, which also contains a report of similar location awareness in the brains of bats.
Hippocampal place cells light up only for certain locations in the environment, and some of these cells clearly preferred the location of the other rat. Besides activation to places, the timing of neural activity in the hippocampus is also important. A ‘refresh rate’ of around 8 Hz dictates how often neurons update their activity, a phenomenon called theta-cycle phase precession. Three-quarters of cells updated based on the other’s location, not only the self’s. “It is very interesting that the trajectories of the other, that is, the past, current, and future positions of the observed rat, are compressively represented in 100-millisecond cycles in the hippocampus,” says Shigeyoshi Fujisawa, research group leader at the RIKEN Brain Science Institute.
In two versions of the maze, the self-rat had to learn to visit the same T-arm as the other or the opposite arm. The majority of neurons were ‘goal-focused’, signaling the location of the goal, but a small number of neurons preferred the other’s location regardless of whether it was on the same side as the goal. There were also cells that fired when either the other or the self were in specific spatial locations. “These cells are not confused,” says Fujisawa. “We can reconstruct the paths of the pair of rats and reliably decode the location of the self or the other from the activity of these joint place cells.”
Fujisawa and colleagues propose that the hippocampus has four different kinds of spatial models, one for the locations of the self, one for that of others, another for joint locations that are tuned relative to where the self and other are, and one for 'common' locations that activate when either the self or other are there (see image). This extends the existing cognitive map theory of how the hippocampus processes spatial locations and memories. “We think the cognitive map in the hippocampus is not just for knowing where the self is located,” says Fujisawa, “but also for plotting the locations of other people, animals, or objects, and to comprehend the spatial environment surrounding the self.”
Reference
- Teruko Danjo, Taro Toyoizumi, Shigeyoshi Fujisawa, "Spatial representations of self and other in the hippocampus", Science, doi: 10.1126/science.aao3898
Contact
Laboratory Head
Shigeyoshi Fujisawa
Laboratory for Systems Neurophysiology
RIKEN Brain Science Institute
Laboratory Head
Taro Toyoizumi
Laboratory for Neural Computation and Adaptation
RIKEN Brain Science Institute
Adam Phillips
RIKEN International Affairs Division
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: adam.phillips@riken.jp
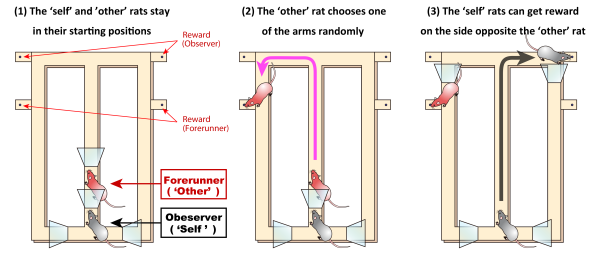
Schematic design of the maze task
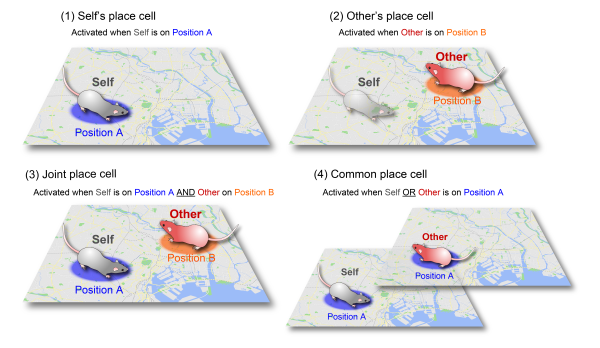
The four types of spatial models for the hippocampus that are proposed in the paper.
