Apr. 5, 2019 Research Highlight Biology
Mapping RNA on a single-cell level
A new analytic method can now comprehensively profile RNA transcripts at single-cell resolution
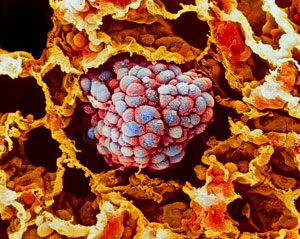 By using their new microfluidic platform to profile RNA at single-cell resolution, RIKEN researchers have shown that different subgroups of lung cancer cells (clump of cells in the center of micrograph) respond differently to a growth factor thought to promote cancer aggression. © Science Photo Library - MOREDUN ANIMAL HEALTH LTD
By using their new microfluidic platform to profile RNA at single-cell resolution, RIKEN researchers have shown that different subgroups of lung cancer cells (clump of cells in the center of micrograph) respond differently to a growth factor thought to promote cancer aggression. © Science Photo Library - MOREDUN ANIMAL HEALTH LTD
A microfluidic technique developed at RIKEN allows scientists to systematically profile RNA transcripts at the single-cell level1. To demonstrate its potential, RIKEN researchers have used the platform to gain insights into how lung cancer cells respond to a growth factor.
The method builds on a procedure called cap analysis gene expression (CAGE) developed more than 15 years ago by Piero Carninci of the RIKEN Center for Integrative Medical Sciences. Unlike most other approaches for characterizing RNA profiles, which start from the end of the transcript and read backwards, CAGE works from the front of the transcript. This enables scientists both to determine the start sites and detect a greater diversity of RNA molecules—not just those that encode proteins, but also enhancers and other regulatory elements.
Jay Shin and Charles Plessy of the RIKEN Center for Integrative Medical Sciences have now adapted CAGE to work at a single-cell resolution. They used a commercially available microfluidics platform called C1, which isolates, processes and prepares individual cells for genetic analysis. Working in collaboration with Carninci, Shin and Plessy then integrated his RNA expression analysis system to create C1 CAGE.
To demonstrate the power of the technology, the team measured RNAs in individual lung cancer cells exposed to a growth factor called TGF-β. This growth factor is generally thought to promote the invasiveness and spread of tumors.
But 24 hours after treatment, the C1 CAGE analysis showed that only one subpopulation of cells expressed genes associated with molecular aggressiveness. Another large population of cells exhibited a more delayed response. “This suggests that there is an asynchronous progression upon TGF-β stimulation,” says Shin.
The RNA profiling also revealed that individual cells display coordinated bursts of expression of RNA from both promoters, the main region of RNA production, and enhancers, subregions that further spur the activity of promoters. No one had ever seen that before at the single-cell resolution, says Shin.
The finding has therapeutic relevance. “It suggests that enhancer RNAs are regulated by a similar machinery as promoters,” Shin says. “This could potentially be manipulated or controlled with anticancer drugs designed to tamp down expression of growth-promoting genes.”
The team compared the new method with a standard protocol for single-cell RNA profiling. “Our C1 CAGE was more accurate, more sensitive, and detected more genes,” says Plessy. Now at the Okinawa Institute of Science and Technology Graduate University, Plessy continues to collaborate with his former RIKEN colleagues to further improve the technology.
“Such optimization will help us detect more transcripts, so that we can better map the promoter and enhancer landscape across different cell types,” says Shin.
Related contents
References
- 1. Kouno, T., Moody, J., Kwon, A. T., Shibayama, Y., Kato, S., Huang, Y., Böttcher, M., Motakis, E., Mendez, M., Severin, J. et al. C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution. Nature Communications 10, 360 (2019). doi: 10.1038/s41467-018-08126-5
