Aug. 11, 2006 Research Highlight Biology Chemistry
Color and light to illuminate biochemical pathways
A new fluorescent label for proteins to reveal greater molecular detail of biochemical pathways
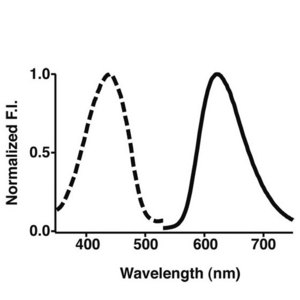 Figure 1: Absorption (dotted line) and emission (full line) spectra of Keima, the new fluorescent protein label developed at the RIKEN Brain Science Institute.
Figure 1: Absorption (dotted line) and emission (full line) spectra of Keima, the new fluorescent protein label developed at the RIKEN Brain Science Institute.
A team of researchers from RIKEN and Hokkaido University have developed a fluorescent label for proteins with a large shift between the colors of light it absorbs and emits (Fig. 1). Their work should allow protein-protein interactions to be tracked more easily, thus revealing biochemical pathways in greater molecular detail.
The new tag can be used in conjunction with existing fluorescent labels in which the color shift is much smaller in the sensitive technique known as fluorescence cross-correlation spectroscopy. The idea is that while both labels can be excited by a single laser, each will fluoresce a different colour. So, two molecular species can be tracked simultaneously.
Fluorescent compounds absorb light at the specific wavelengths, or colors, which provide enough energy to boost electrons to higher energy levels. Light is emitted as the electrons fall back to their normal energy levels. But some energy is always lost as heat in the process, and so the wavelength of the light emitted is always longer, or less energetic, than that of the light absorbed. The difference is known as the Stokes shift.
Researchers from RIKEN’s Brain Science Institute in Wako have worked with colleagues the Research Institute for Electronic Science at Hokkaido University in Sapporo to create the new fluorescent tag that has an absorption energy typical of labels already in use, but a Stokes shift larger than usual, so that its fluorescence is easy to distinguish.
 Figure 2: The stony coral Montipora from which the violet-coloured protein used as a basis for Keima was extracted.
Figure 2: The stony coral Montipora from which the violet-coloured protein used as a basis for Keima was extracted.
As reported in Nature Biotechnology 1, the team developed the label from a violet-coloured protein found in the stony coral Montipora (Fig. 2) by changing its amino acid components and side chains at critical points until they generated a family of compounds that exhibited particular characteristics. The researchers have called the fluorescent protein Keima, after a Japanese chess piece that moves in the hopping manner of a knight.
The research team was able to demonstrate Keima’s value by using it to follow protein-protein interactions. For instance, by attaching Keima and a cyan fluorescent protein either side of a protein sequence known to be cleaved by a particular enzyme, the researchers could study details of the interaction as the fragments with different coloured fluorescent tags broke apart. They also tracked another interaction whereby two tagged compounds came together.
“This is a sensitive and quantitative technique for investigating protein-protein interactions,” says team leader Atsushi Miyawaki. “We are developing it to use in many systems. And pharmaceutical companies are interested in using it.”
References
- 1. Kogure, T., Karasawa, S., Araki, T., Saito, K., Kinjo, M. & Miyawaki, A. A fluorescent variant of a protein from the stony coral Montipora facilitates dual-color single-laser fluorescence cross correlation spectroscopy. Nature Biotechnology advance online publication, 30 April 2006. doi: 10.1038/nbt1207
