Nov. 22, 2007 Research Highlight Biology
Thinking outside the cell
A recently developed experimental system provides new insight into how tiny RNA molecules keep a rein on gene activity
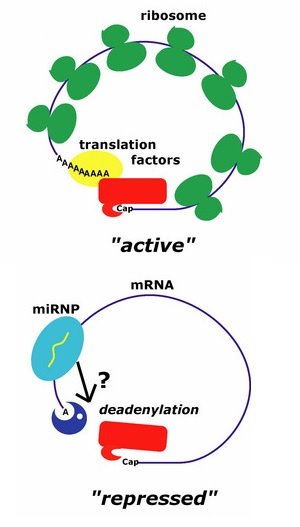 Figure 1: A model for how let-7 represses translation. Under normal conditions (upper), the cap and polyadenosine tail of an mRNA are bound by proteins (red and yellow, respectively) that make translation possible. When the miRNP is bound (lower), its component proteins trigger transcript deadenylation, preventing translation.
Figure 1: A model for how let-7 represses translation. Under normal conditions (upper), the cap and polyadenosine tail of an mRNA are bound by proteins (red and yellow, respectively) that make translation possible. When the miRNP is bound (lower), its component proteins trigger transcript deadenylation, preventing translation.
Every gene’s activity is transmitted by messenger RNA (mRNA) transcripts, which subsequently serve as the template for protein-building—a process known as translation. Biologists have historically underestimated the humble mRNA, viewing it as little more than the short-lived intermediate between gene and protein, but groundbreaking work over the last decade or so has considerably changed that view.
MicroRNAs are a recently discovered class of tiny RNA transcripts that don’t encode proteins at all, but instead act as regulators of the activity of other genes. These microRNAs partner with cellular proteins to form the microRNA–protein complex (miRNP). The RNA sequences of this complex enable it to directly bind to partially complementary sequences on target mRNAs. This binding can lead to greatly reduced protein production, although the mechanism of this process remains unclear.
Motoaki Wakiyama, a senior scientist in Shigeyuki Yokoyama’s laboratory at the RIKEN Genomic Sciences Center in Yokohama, found it difficult to study this process in the complex environment of the living cell, and so the group attempted to instead recreate microRNA activity in a test tube. “In a cell-free system, you can specifically focus on translational control,” says Yokoyama, “and it is very easy to handle various reaction conditions.”
Yokoyama’s group mixed extracts from mammalian cell lines expressing various miRNP component proteins in order to understand the mechanism of the microRNA let-7 1. After identifying conditions that recreate let-7 repression of target transcripts, the group focused on exactly how the microRNA blocks translation. They found that this repression relies on interactions with two core structural components of mRNA—a chemical ‘cap’ at one end, and a long ‘tail’ of adenosine nucleotides at the other. In fact, miRNP binding is followed by dramatic shortening of this tail—a process known as deadenylation—which in turn blocks mRNA translation (Fig. 1).
“The most surprising finding is that microRNA induces deadenylation, causing translational repression but not rapid mRNA degradation,” says Yokoyama. Based on current evidence, he believes this may represent a general mechanism by which microRNAs down-regulate target genes. “It is known that polyadenylation and deadenylation control mRNA translation,” he says.
These findings lay important groundwork for future microRNA studies, and Yokoyama is now exploring the functional roles of individual miRNP components, but he also emphasizes the value of the experimental model developed here. “The recapitulation of microRNA-mediated translational repression in a test tube provides a powerful tool for studying microRNA function,” says Yokoyama.
References
- 1. Wakiyama, M., Takimoto, K., Ohara, O. & Yokoyama, S. Let-7 microRNA-mediated mRNA deadenylation and translational repression in a mammalian cell-free system. Genes & Development 21, 1857–1862 (2007). doi: 10.1101/gad.1566707
