Apr. 19, 2013 Research Highlight Biology
Asymmetry makes the protein pump go round
First high-resolution structures provide clues to the mechanism of an important molecular rotary motor
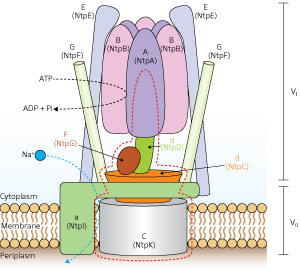 Figure 1: Schematic model of V-ATPase in Enterococcus hirae, showing the A3B3 barrel complex (top center) and the DF complex (immediately below). © 2013 Takeshi Murata et al., RIKEN Systems and Structural Biology Center
Figure 1: Schematic model of V-ATPase in Enterococcus hirae, showing the A3B3 barrel complex (top center) and the DF complex (immediately below). © 2013 Takeshi Murata et al., RIKEN Systems and Structural Biology Center
The protein vacuolar ATPase (V-ATPase) is an attractive drug target due to its crucial role in several disease conditions including osteoporosis and the spread of cancer. It is found within the membranes of many organelles, where it pumps protons from one side of the membrane to the other. A team of researchers led by Takeshi Murata and colleagues from the RIKEN Systems and Structural Biology Center has now produced the first high-resolution structures of a bacterial version of this important protein, revealing previously unknown details of its pumping mechanism1.
V-ATPase in the bacterium Enterococcus hirae is closely related to its human equivalent, consisting of nine different protein-chain subunits. One part of the protein, V0, is hydrophobic and sits in the membrane, while the other part, V1, is hydrophilic and juts out into the cell body. As a result of previous work in which a team led by Murata produced, purified and assembled the protein subunits comprising V1 in the laboratory, it was revealed that V1 consists of a hexagonal barrel-like complex built from three catalytic ‘A’ subunits alternating with three non-catalytic ‘B’ subunits (Fig. 1). Another complex, known as DF, fits into the center of the barrel and rotates.
In their most recent work, Murata’s team solved x-ray crystallography structures for the A3B3 complex, as well as for the A3B3 complex attached to the DF complex and in association with an analog of ATP—a biological energy carrier. Much to their surprise, they found that although the three A chains all had the same sequence, as did the three B chains, the A3B3 complex fitted together in an asymmetric fashion.
The A3B3 complex should contain three pockets between the A and B subunits, into which ATP can bind. When the ATP analog was added, however, the researchers found it bound in only two of the pockets because of the asymmetry. Adding the DF complex increased the asymmetry, causing the three pockets to develop different morphologies—particularly with respect to positioning of a projection involving the amino acid arginine, which assists the breaking of an ATP phosphate bond to release the energy that powers the rotary ion pump. They also found that the shape of the complex appears to change during rotation.
“This structural study will be useful for designing future drugs targeting V-ATPase,” says Murata. “We now want to perform further experiments to understand the rotational mechanism more precisely.”
References
- 1. Arai, S., Saijo, S., Suzuki, K., Mizutani, K., Kakinuma, Y., Ishizuka-Katsura, Y., Ohsawa, N., Terada, T., Shirouzu, M., Yokoyama, S.et al.Rotation mechanism of Enterococcus hirae V1-ATPase based on asymmetric crystal structures. Nature 493, 703–707 (2013). doi: 10.1038/nature11778
