Aug. 15, 2014 Research Highlight Biology
Deflecting protein production
Insights into the mechanistic details of protein synthesis could inform efforts to manipulate the genetic code
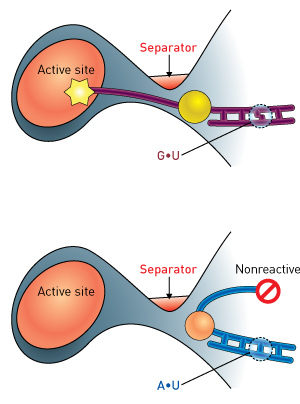 Figure 1: In the naturally occurring guanine–uracil ‘identity set’ pairing (top), the tRNA acceptor region (yellow) is properly positioned at the enzyme’s active site for chemical modification with the amino acid alanine. In the adenine–uracil mutant (bottom), the tRNA is deflected away from the active site and alanine addition is blocked. Reproduced, with permission, from Ref. 1 © 2014 M. Naganuma et al.
Figure 1: In the naturally occurring guanine–uracil ‘identity set’ pairing (top), the tRNA acceptor region (yellow) is properly positioned at the enzyme’s active site for chemical modification with the amino acid alanine. In the adenine–uracil mutant (bottom), the tRNA is deflected away from the active site and alanine addition is blocked. Reproduced, with permission, from Ref. 1 © 2014 M. Naganuma et al.
The recipe for any given protein is written out as a series of ‘codons’, each of which encodes a particular amino acid. These amino acids are delivered via transfer RNA (tRNA) molecules, which feature an ‘anticodon’ element that recognizes a particular codon. For this system to function properly, every tRNA must be linked to the correct amino acid—a process that is mediated by a family of enzymes called aminoacyl-tRNA synthetases. A research team led by Shigeyuki Yokoyama from the RIKEN Structural Biology Laboratory and Paul Schimmel of The Scripps Research Institute in the United States has now learned how one such enzyme achieves such selective coupling1.
Decades ago, Schimmel discovered a single base-pair within the tRNA associated with the amino acid alanine (tRNAAla) that serves as an ’identity set’ critical for recognition and processing by its associated aminoacyl-tRNA synthetase enzyme. Strangely, this base-pair resides far from the active site of the enzyme that actually attaches alanine to the tRNA. “It was a complete mystery how these nucleotides indirectly affect the active site of the enzyme,” says Yokoyama. “Furthermore, it seemed almost impossible that a tRNA could depend so strictly on a single base-pair interaction.”
Yokoyama therefore set about analyzing the structure of complexes formed by two different versions of tRNAAla and its aminoacyl-tRNA synthetase. In general, the nucleobase adenine preferentially pairs with the nucleobase uracil. However, the tRNAAla identity set consists of a less common ‘wobble pairing’ between the nucleobases uracil and guanine. The introduction of a mutation that changed this naturally occurring guanine–uracil pairing to adenine–uracil had subtle but important consequences. The nucleobases in each pairing bind with different geometries, and the mutation introduces a curvature in the tRNA structure (Fig. 1). The curvature deflects the acceptor region of the tRNA away from the active site of the enzyme such that the nucleotides that would normally be joined to alanine are instead physically masked by a ‘separator’ segment of the enzyme. This single mutation was sufficient to dramatically reduce the efficiency of this enzyme-mediated reaction.
Yokoyama’s group is actively engaged in manipulating the interactions between tRNAs and synthetase enzymes as a means of producing proteins that incorporate novel amino acids with distinctive functional properties, and Yokoyama now hopes to learn whether similar principles apply in other tRNA–aminoacyl-tRNA synthetase pairings. “These findings could open the door for dramatic developments in the expansion of the genetic code,” he says.
References
- 1. Naganuma, M., Sekine, S., Chong, Y. E., Guo, M., Yang, X.-L., Gamper, H., Hou, Y.-M., Schimmel, P. & Yokoyama, S. The selective tRNA aminoacylation mechanism based on a single G•U pair. Nature 510, 507–511 (2014). doi: 10.1038/nature13440
