Feb. 10, 2017 Research Highlight Biology
Unraveling the complexity of cellular reactions
A mathematical model that captures the structure of chemical reaction networks in biological systems could advance the study of cellular functions
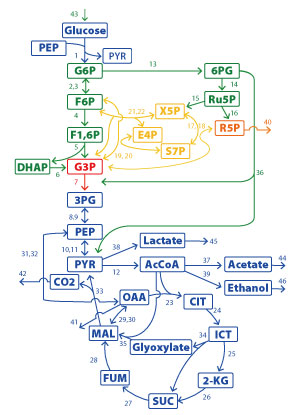 Figure 1: An example of a chemical reaction network in the bacterium Escherichia coli. Reprinted with permission from Ref. 1. Copyright (2017) by the American Physical Society.
Figure 1: An example of a chemical reaction network in the bacterium Escherichia coli. Reprinted with permission from Ref. 1. Copyright (2017) by the American Physical Society.
The responses of chemical reaction systems in living cells to changes in the activities and abundances of enzymes can be predicted using information of the chemical network structure alone, two RIKEN researchers have found1. By constructing an elegant mathematical method known as structural sensitivity analysis, the pair has shown that it is possible to investigate cellular functions without having detailed data on the chemical reactions themselves—a finding that opens new possibilities for the study of disease and drug discovery.
Life is the product of a complex cascade of interrelated chemical and physicochemical interactions within and between cells. These interactions are mediated and modulated by enzymes and their metabolites. Consequently, changes in the activity and abundance of a single enzyme can lead to major alterations in biological function. Researchers often study such changes as potential disease mechanisms and targets for possible therapies. However, obtaining quantitative information on the activity and abundance of enzymes and all of the biomolecules involved in the reactions is an extremely daunting task.
“The chemical reaction network is dynamic and involves many reactions that are connected by sharing products and substrates with other reactions,” explains Atsushi Mochizuki of the RIKEN Theoretical Biology Laboratory, who worked with colleague Takashi Okada on the study. “However, our understanding of the dynamic behavior of such networks is poor due to their complexity. Our study provides a clear and strong mathematical solution to this general biological problem.”
Mochizuki and Okada developed a mathematical structural sensitivity analysis scheme that models how much of the network is affected by a change in that activity or abundance of a particular enzyme. Despite its simplicity—it involves just ordinary differential equations and reaction stoichiometry—the model directly connects the network typology to system response and has revealed new levels of structure in what are well-studied systems. The theorem is also mathematically beautiful.
Furthermore, the study has revealed that the response patterns, that is the distribution of nonzero responses of chemical concentrations in the network, generally exhibit two characteristic patterns—localization and hierarchy—and that the nonzero response patterns can be completely explained by a new theorem, which the researchers call the law of localization.
Mochizuki and Okada demonstrate their method by applying it to several hypothetical and real-life chemical reaction networks, including the metabolic network of the bacterium Escherichia coli (Fig. 1).
The mathematical scheme promises to be a powerful tool for gleaning vital information about the complexities of the chemical reactions that occur in cells.
Related contents
- iTHES: unraveling multi-scale problems through interdisciplinary theory
- Plant enzymes reveal complex secrets
References
- 1. Okada, T. & Mochizuki, A. Law of localization in chemical reaction networks. Physical Review Letters 117, 048101 (2016). doi: 10.1103/PhysRevLett.117.048101
