Jul. 10, 2015 Research Highlight Biology
Uncovering the secrets of antibacterial response
New insights into a well-known immune pathway could help researchers fight tuberculosis and other infectious diseases
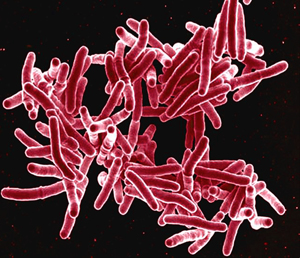 Figure 1: The bacterium Mycobacterium tuberculosis, responsible for tuberculosis, can trigger a cell signaling response in macrophages that elicits a broader inflammatory response. Courtesy: National Institute of Allergy and Infectious Diseases
Figure 1: The bacterium Mycobacterium tuberculosis, responsible for tuberculosis, can trigger a cell signaling response in macrophages that elicits a broader inflammatory response. Courtesy: National Institute of Allergy and Infectious Diseases
A protein with a previously unrecognized—but apparently critical—role in a well-studied immune pathway has been identified by a team jointly led by Harukazu Suzuki of the RIKEN Center for Life Science Technologies in Japan and Frank Brombacher in South Africa1.
Immune cells known as macrophages fend off pathogenic bacteria by destroying infected cells. This process triggers a signaling cascade known as the classical activation pathway, which in turn stimulates a broader inflammatory response. However, despite extensive study, many of the molecular mechanisms that govern the complex transcriptional regulation in classically activated macrophages are not well understood.
Since the protein interferon-γ (IFN-γ) helps initiate classical activation, the researchers initially set out to identify genes switched on by IFN-γ. They observed particularly strong activity from Batf2, which expresses a protein belonging to a family of transcription factors that regulate genes but are otherwise poorly characterized. The scientists found that they could interfere with many aspects of IFN-γ-induced classical activation by experimentally manipulating macrophages to reduce Batf2 activity.
The bacterium responsible for tuberculosis, Mycobacterium tuberculosis (Fig. 1), can also trigger classical activation, and the researchers observed an equally prominent role for Batf2 in this response. Indeed, many genes activated by the Batf2 protein make important contributions to host defense and to promoting inflammation, providing further evidence that the gene is a valuable mediator of classical activation.
Although the Batf2 protein can switch genes on and off, it lacks the structural components necessary to bind to DNA. Consequently, it has to rely on a DNA-binding ‘partner’. By examining the sequences of known Batf2 target genes, Suzuki and his co-workers zeroed in on IFN regulatory factor-1 (Irf1) as the most likely collaborator. ‘Knockdown’ experiments that reduced macrophage production of Irf1 confirmed this, with many of the same changes in gene expression observed as in the Batf2-deficient cells.
This discovery of a major component of the macrophage response that had gone unnoticed for so long surprised Suzuki. “This response has been extensively analyzed by many groups,” he says. “I initially thought that Batf2 might be a minor modulator of classical activation, but I now believe that Batf2 plays a central role, together with Irf1.”
Suzuki’s group intends to delve deeper into the nuts and bolts of the machinery for macrophage activation. Suzuki is also working with Brombacher to learn whether this pathway might offer a target for stimulating the immune response against tuberculosis. “We are currently analyzing the in vivo effect of Batf2 on M. tuberculosis infection in mice,” he says.
References
- 1. Roy, S., Guler, R., Parihar, S. P., Schmeier, S., Kaczkowski, B., Nishimura, H., Shin, J. W., Negishi, Y., Ozturk, M., Hurdayal, R. et al. Batf2/Irf1 induces inflammatory responses in classically activated macrophages, lipopolysaccharides, and mycobacterial infection. The Journal of Immunology 194, 6035–6044 (2015). doi: 10.4049/jimmunol.1402521
