May 28, 2010 Press Release Biology
Molecular circuit uncovered governing key cell-fate decisions
Critical missing links in a signaling-transcription cascade responsible for pivotal cell-fate decisions have been described for the first time in a paper in Cell. Identified through a combination of simulations and experiments, the links are part of circuit-like molecular control mechanisms for converting analog signals into binary responses central to the development of all cells.
The question of how identical cells develop into distinct cell types using the same signaling pathways is integral to our understanding of the cell life cycle. The mechanisms that determine cell fate decisions, leading cells with the same genes to distinct developmental outcomes, remain however poorly understood.
To study cell fate decisions, a research team headed by scientists at the RIKEN Research Center for Allergy and Immunology (RCAI) and the University College Dublin administered growth factors to MCF-7 breast cancer cells and analyzed responses in the extracellular regulated kinase 1/2 (ERK) cascade. Whereas one growth factor (epidermal growth factor or EGF) induces transient ERK activity leading to cell proliferation, the other (heregulin or HRG) induces ERK activity that is sustained, triggering cell differentiation. Connecting these analog ERK signaling patterns to their cell fates (proliferation/differentiation) is the phosphorylated transcription factor c-Fos, whose digital all-or-none expression acts as the output of the signaling system.
Comparing observational data with results of mathematical simulations, the researchers arrived at a "molecular circuit" model for c-Fos mediated cell differentiation composed of negative feedback loops, feed-forward loops and logical AND gates that reduce noise and generate stable output signals. The discovery of these simple circuit components, which are believed to govern differentiation across a variety of different cell types, provides fundamental insights into the underlying logic of cell-fate decision processes, opening the door to applications in areas such as regenerative medicine.
Contact
Mariko Okada-Hatakeyama
Laboratory for Cellular Systems Modeling
RIKEN Research Center for Allergy and Immunology (RCAI)
Tel: +81-(0)45-503-9302 / Fax: +81-(0)45-503-9613
Jens Wilkinson
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr@riken.jp
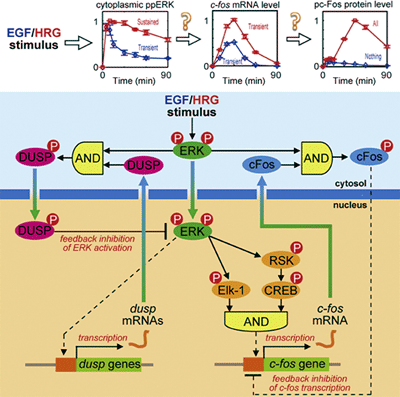
Figure 1: Graphical abstract of molecular circuit.
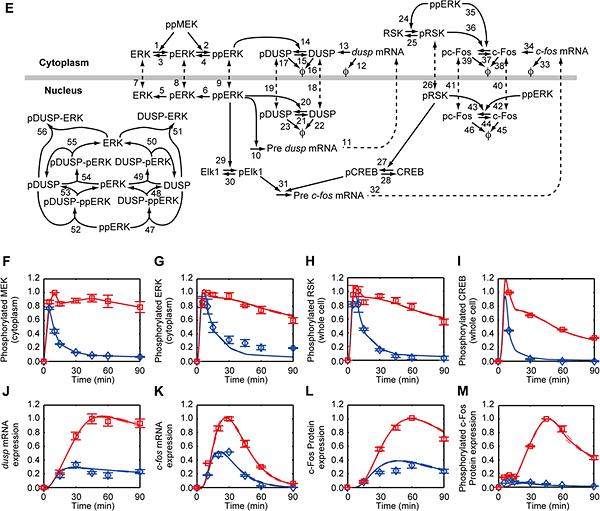
Figure 2: Quantitative time-course analysis of c-Fos expression network
Responses to EGF (proliferation signal; red) and HRG (differentiation signal; blue) and model's schematic.
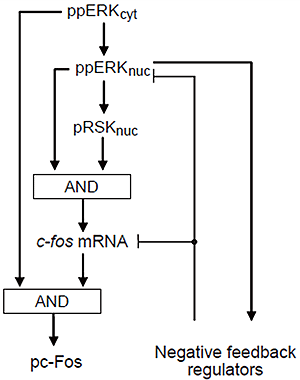
Figure 3: Control theory embedded in the model
AND gate, feedforward and feedback regulation induce stable binary response of phosphorylated c-Fos protein (pc-Fos).
