Sep. 9, 2014 Press Release Biology
Broken signals lead to neurodegeneration
Researchers from the RIKEN Brain Science Institute in Japan, in collaboration with Juntendo University and the Japan Science and Technology Agency, have discovered that a cell receptor widely involved in intracellular calcium signaling—the IP3R receptor—can be locked into a closed state by enzyme action, and that this locking may potentially play a role in the reduction of neuron signaling seen in neurodegenerative diseases such as Huntington's and Alzheimer's disease.
In the research published today in the Proceedings of the National Academy of Sciences, the scientists reported experiments in human cells and a mouse model of Huntington's disease revealing that transglutaminase type 2—a protein cross-linking enzyme elevated in the cells of patients with neurodegenerative diseases—interacts with the IP3R receptor to lock it in a closed non-functional conformation preventing it from fulfilling its essential calcium-releasing role. They identified a specific amino acid site on the receptor, Gln2746, where the modification takes place, deepening our understanding of how receptors are locked and potentially opening the door to studies on other functional proteins that are also regulated by conformational changes.
The IP3R channel, which is located in the endoplasmic reticulum, a protein assembly and transport compartment, plays a crucial role in intracellular calcium signaling, and is involved in a wide range of cell functions including mitochondrial energy production and the regulation of autophagy, the process through which cells consume and degrade unused components to maintain a healthy balance of functional proteins. Although autophagy is normally a mechanism that sustains cell maintenance, it can also trigger a loss of cell function and has been associated with prominent diseases including Huntington's disease, Alzheimer's disease, and Parkinson's disease.
In this work, the scientists propose a general model under which abnormal IP3R-mediated calcium signaling caused by the action of transglutamase type 2 leads to cellular dysfunction and subsequently to the emergence of progressive brain dysfunction. Transglutaminase 2 activation is commonly associated with inflammation and stress, and its action on the IP3R channel might provide an explanation for the initiation and progression steps common to different neurodegenerative diseases.
According to Katsuhiko Mikoshiba, who led the study, "We think that the mechanism we identified in this study could provide us with a more general model of other diseases both of the brain and other parts of the body, where transglutaminase type 2 is upregulated. We hope that this insight could eventually lead to the development of new drug therapies for a number of neurodegenerative diseases that place a high burden on patients and society."
Reference
- Kozo Hamada, Akiko Terauchi, Kyoko Nakamura, Takayasu Higo, Nobuyuki Nukina, Nagisa Matsumoto, Chihiro Hisatsune, Takeshi Nakamura, and Katsuhiko Mikoshiba, Aberrant calcium signaling by transglutaminase-mediated posttranslational modification of inositol 1,4,5-trisphosphate receptors, PNAS, doi: 10.1073/pnas.1409730111
Contact
Laboratory Head
Katsuhiko Mikoshiba
Laboratory for Developmental Neurobiology
RIKEN Brain Science Institute
Jens Wilkinson
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr@riken.jp
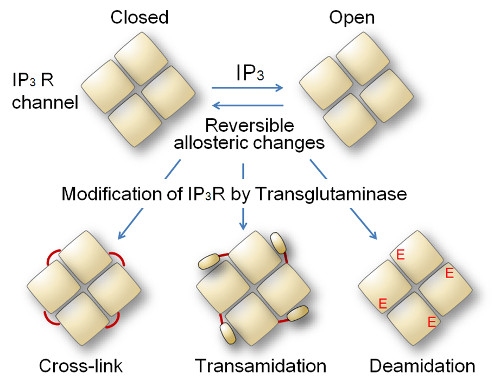
Model for transglutaminase-mediated modification of IP3R
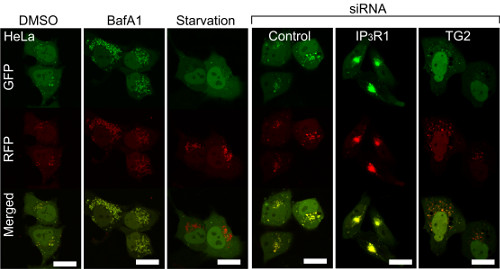
Deranged autophagy by IP3R and transglutaminase regulation
