Feb. 13, 2015 Press Release Biology
FANTOM5 project discovers general rules governing how cells change
Using a comprehensive analysis of RNA expression in different cell types, scientists from the RIKEN-led FANTOM5 consortium have made major strides toward resolving an outstanding mystery in biology. In the work, published in Science, they showed that when cells undergo phenotype changes such as differentiation into specialized cell types, the initial activation takes place at DNA regions called enhancers, a type of regulatory "switch" which are typically located far from the genes that they activate.
It is well known that cells undergo tightly controlled patterns of transcription changes—modifying which genes are expressed, and at what level—both during development, when they change to specialized cell types, and in response to stresses or infection. It was understood that both promoters—DNA regions located near the genes they regulate—and enhancers, which are located far away, played a role, but the sequence of events was not well understood, and it was generally assumed that enhancers and promoters acted more or less simultaneously.
The current research, which examined 19 human cell types and 14 mouse cell types undergoing a variety of cellular changes, showed that it is the activation of enhancers that triggers the coordinated waves of change that end up dramatically changing the phenotypes of the cells. In particular, enhancers are activated in the first 15 minutes after stimuli, and then activate a specific type of regulatory gene (transcription factors) at 30-100 minutes, which in turn have the ability to activate other genes over time, forming a cascade of changes. These patterns were shared across all the cells studied, but the individual genes and enhancers that made up the patterns were specific to each cell type. This suggests that the team has uncovered underlying rules for how cells differentiate, and that these rules apply to all mammalian cells. This knowledge is an important part of the puzzle in understanding how cells and ultimately human bodies work in health and disease.
The group used CAGE technology, a method developed in the RIKEN-led FANTOM project to identify the starting points of transcription sites, in a variety of time courses including stem cells, cells that were part-way on the path to terminal differentiation, and differentiated cells responding to stimuli such as growth factors or pathogens.
This is another important milestone for the FANTOM consortium, which in March 2014 used CAGE to identify the promoters and enhancers that are used in most existing mammalian cell types. The project is now using CAGE technology to identify the sequence of events through which cells change.
According to Erik Arner of the RIKEN Center for Life Science Technologies (CLST), the first author, "This is a very exciting finding as it has given us a better understanding of the complex interplay that takes place as cells adapt and differentiate".
Piero Carninci, Deputy Director of the RIKEN Center for Life Science Technologies, says, "These results are an important example of how a large international collaboration, in this case comprising 114 institutions in more than 20 countries, can contribute to our understanding of fundamental questions in biology by unveiling the basic rules of biological regulation. We are proud that 15 years from its inception, the focus and methodologies of the FANTOM consortium are still evolving to remain at the forefront of genomics".
And looking at the implications, Yoshihide Hayashizaki, program director of the RIKEN Preventive Medicine & Diagnosis Innovation Program, says, "Understanding the leading role of enhancers could lead to improved methods for generating differentiated cells from ES or iPS cells. Furthermore, since it induces tissue-specific gene expression and perhaps epigenomic alternation, enhancer activity could be a key to achieving effective cell manipulation".
Reference
- Erik Arner et al., Transcribed enhancers lead waves of coordinated transcription in transitioning mammalian cells, Science, doi:10.1126/science.1259418
Contact
Team Leader
Piero Carninci
Erik Arner
Genome Information Analysis Team
Life Science Accelerator Technology Group
Division of Genomic Technologies
RIKEN Center for Life Science Technologies
Program Director
Yoshihide Hayashizaki
RIKEN Preventive Medicine & Diagnosis Innovation Program
Adam Phillips
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr [at] riken.jp
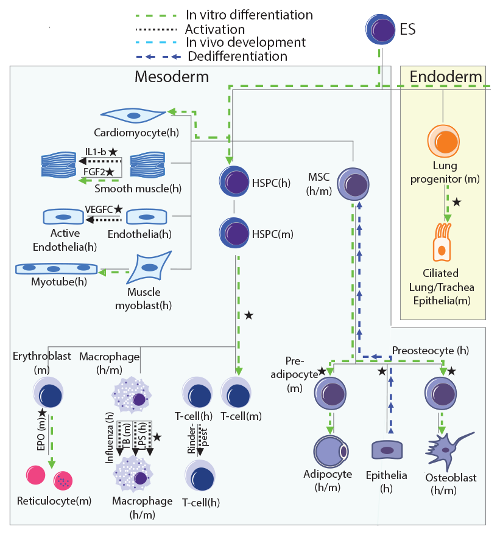
Illustration of the time course experiments included in the study, arranged following a development tree
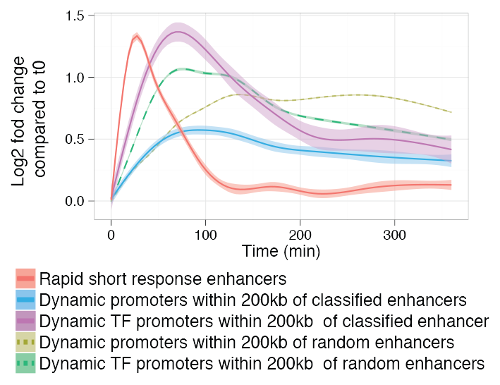
Smoothed mean expression over time for all enhancers classified into the rapid short response group and all differentially expressed proximal (+/-200kb) promoters, split by gene type