Jul. 22, 2011 Research Highlight Biology
Silence of the genes
As in other multicellular organisms, plants have evolved mechanisms to maintain genome stability and integrity
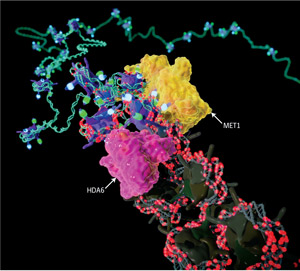 Figure 1: In Arabidopsis chromatin, the enzymes HDA6 and MET1 convert epigenetic information to make it ‘silent’. Reproduced with permission © 2011 CYCLOID Inc. All rights reserved
Figure 1: In Arabidopsis chromatin, the enzymes HDA6 and MET1 convert epigenetic information to make it ‘silent’. Reproduced with permission © 2011 CYCLOID Inc. All rights reserved
A molecular mechanism by which gene silencing is regulated at the genome-wide level in plants has been uncovered by a research team led by Motoaki Seki of the RIKEN Plant Science Center, Yokohama1. The researchers propose that a similar mechanism may also help to protect plant genomes from the potentially harmful effects of DNA elements, such as transposons, or ‘jumping genes’. “If left unhindered, transposable elements can cause havoc in the genome, for example by inserting themselves into essential genes,” says Seki.
The DNA of eukaryotes—organisms with nucleated cells—is packaged in a complex structure called chromatin within chromosomes. Chromatin also contains DNA-binding proteins called histones. When in its open conformation, known as euchromatin, the DNA is accessible to transcription factors, allowing gene expression to proceed. However, when in its highly condensed form—heterochromatin—gene expression is silenced.
The transition from euchromatin to heterochromatin requires chemical modification of both DNA and histones. These so-called epigenetic changes involve the methylation of DNA by enzymes called DNA methyltransferases, and the elimination of epigenetic marks on histones by other enzymes called histone deacetylases. In addition to silencing gene expression, heterochromatin formation may protect against the potentially damaging effects of transposons by blocking their replication.
Seki and his colleagues studied the regulation of heterochromatin formation in Arabidopsis thaliana, a small flowering related to the mustard plant. “Arabidopsis is a widely used model species for studying epigenetic changes in plants,” explains Seki.
Uniquely in plants, DNA methylation resulting in heterochromatin formation is triggered by small RNA molecules. This process is known as RNA-directed DNA methylation, and involves the DNA methyltransferase MET1 and the histone deacetylase HDA6 (Fig. 1). However, the overall role of HDA6 in heterochromatin formation remained unclear.
By comparing the RNA transcript profiles of normal and mutant plants lacking functional HDA6, the researchers identified 157 target genes spread across the Arabidopsis genome. In some target genes in the mutant plants they found that DNA methylation was completely lost, allowing these genes to be expressed. They also found that the target specificity of HDA6 was unexpectedly much greater than that of MET1.
“Our findings suggest that HDA6 recruits MET1 to specific target genes, allowing it to regulate gene silencing on a genome-wide scale,” says Seki.
In addition to this general role, the researchers propose that HDA6 may regulate transposon silencing through heterochromatin formation in plant gametes. They also express the hope that their research will help illuminate related processes in humans.
References
- 1. To, T.K., Kim, J.-M., Matsui, A., Kurihara, Y., Morosawa, T., Ishida, J., Tanaka, M., Endo, T., Kakutani, T., Toyoda, T., et al. Arabidopsis HDA6 regulates locus-directed heterochromatin silencing in cooperation with MET1. PLoS Genetics 7, e1002055 (2011). doi: 10.1371/journal.pgen.1002055
