Aug. 3, 2012 Research Highlight Medicine / Disease
Uncovering secrets of a childhood disease
In-depth genomic analysis helps researchers inch closer to an understanding of an enigmatic cardiovascular disorder that predominantly affects young East Asian children
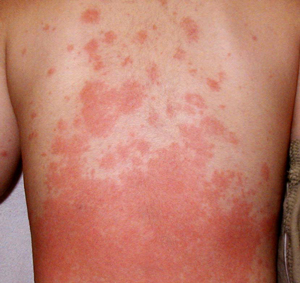 Figure 1: Kawasaki disease may be diagnosed based on inflammatory symptoms, such as the rash shown here. © 2012 Yoshihiro Onouchi, RIKEN Center for Genomic Medicine, Yokohama
Figure 1: Kawasaki disease may be diagnosed based on inflammatory symptoms, such as the rash shown here. © 2012 Yoshihiro Onouchi, RIKEN Center for Genomic Medicine, Yokohama
For more than 40 years now, scientists have grappled with the mysteries of Kawasaki disease (KD). New genetic data from a large team of Japanese and American scientists may help fill in the gaps1. The condition (Fig. 1), which primarily affects children under the age of five, is often harmless. However, up to a quarter of all children with KD will succumb to potentially serious cardiovascular damage if left untreated, including dilation of the coronary artery and aneurysm. “Patients who have giant aneurysms have an increased risk of sudden death, require anticoagulant therapy and have exercise limitations,” explains Yoshihiro Onouchi of the RIKEN Center for Genomic Medicine in Yokohama.
KD is the leading cause of heart disease among children in much of the developed world, but the problem is especially acute in East Asia, particularly Japan. Estimates suggest that in the US and UK, respectively, only 20.8 and 8.4 out of every 100,000 children aged four or younger will be diagnosed in a given year. In contrast, Japan and Korea respectively see 222.9 and 113.1 new cases of KD each year per 100,000 children in this age group.
Experts hypothesize that the condition may be triggered by infection, although the causative agent has not been identified. Nevertheless, there is striking evidence for a hereditary component and Onouchi has been in active pursuit of genetic risk factors for more than a decade. “I am also a pediatrician and started focusing on the genetic aspect of KD in 1997 as a postgraduate student,” says Onouchi. Most recently, he and colleagues led an extensive population genetics study that validated some previously identified candidate risk factors while also spotlighting some promising new ones.
Sifting through sequences
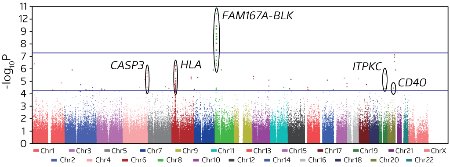 Figure 2: Statistical analysis of SNPs associated with KD revealed a number of ‘hits’ in various chromosomal regions (circled dots), but the strongest association was found for SNPs in the vicinity of the BLK gene. © 2012 NPG
Figure 2: Statistical analysis of SNPs associated with KD revealed a number of ‘hits’ in various chromosomal regions (circled dots), but the strongest association was found for SNPs in the vicinity of the BLK gene. © 2012 NPG
Genome-wide association studies (GWAS) are the weapon of choice for exploring the roots of hereditary disease risk. In a GWAS, researchers perform genomic analyses of large cohorts of patients afflicted with a given condition, analyzing small DNA sequence changes known as single-nucleotide polymorphisms (SNPs) scattered throughout the genome. By statistically determining which SNP variants are disproportionately present in the genomes of KD patients versus demographically similar cohorts of unaffected controls, it becomes possible to identify regions containing risk factors and, in some cases, actual disease-causing mutations.
In this study, Onouchi and colleagues analyzed SNPs from a population of Japanese children, including 447 KD patients and 3,397 controls. They subsequently validated the top candidates from this round by performing confirmatory analyses in other case-control pairs to ensure that the findings were statistically robust.
Among their new discoveries was a clear and strong disease association for several SNPs in the vicinity of the BLK gene (Fig. 2), which encodes a protein associated with signaling by antibody-producing B cells. The researchers also identified a strong association for a SNP in the vicinity of the CD40 gene. CD40 is expressed by a host of different immune cell types, and previous studies have linked variations in this gene with autoimmune and inflammatory disease. These specific variations can markedly alter the function of the encoded CD40 protein and Onouchi and colleagues found preliminary evidence that they may also contribute to the etiology of KD, although further confirmation will be necessary.
This study also spotlighted SNPs near FCGR2A, a gene that has been strongly linked to KD in analyses of both Asian and European cohorts, and which also highlights a potential immune basis for the disease. “The FCGR2A protein plays an important role in up-regulating the inflammatory response,” says Onouchi. “However, there is also a cluster of receptor genes around FCGR2A, any one of which could be a true susceptibility gene.” Two other previously identified KD-associated loci, ITPKC and CASP3, were among the top hits identified in this study.
Filling in the map
This study did not reveal statistically meaningful associations for several other loci identified in previous GWAS, and this current dataset also revealed possible links to genes whose association with KD has not been observed in other cohorts. However, Onouchi notes that this is simply an inherent challenge in performing GWAS—particularly for a condition with such a skewed geographic distribution. “Every GWAS for KD has the limitation of small sample size, such that susceptibility loci with moderate effect size might be missed,” he says, “and it is important to recognize that relative contribution of each susceptibility locus can differ among ethnicities.” He also notes that a recent independent analysis by Taiwanese researchers has validated many of the loci identified both in this study and in previous work.
Collectively, these results may not resolve the mystery of KD, but they provide tantalizing, and somewhat unexpected, insights that could steer future investigations. For example, multiple links between risk factors associated with KD and autoimmune conditions like lupus and rheumatoid arthritis could imply related disease pathways. “The data suggest the possibility that there is some overlap between pathophysiology of these diseases,” says Onouchi. “I also think it is surprising that the same genetic factors play roles in the pathogenesis of different diseases—once in early childhood, and then later in adulthood.”
In follow-up studies, he and his colleagues intend to delve deeper in their genomic analysis of high-risk East Asian populations, with the aim of charting out the disease’s natural history, as well as potential indicators of disease severity that might help guide the treatment of KD patients at greatest risk from their disease.
References
- 1. Onouchi, Y., Ozaki, K., Burns, J.C., Shimizu, C., Terai, M., Hamada, H., Honda, T., Suzuki, H., Suenaga, T., Takeuchi, T. et al. A genome-wide association study identifies three new risk loci for Kawasaki disease. Nature Genetics 44, 517–521 (2012). doi: 10.1038/ng.2220
About the Researcher
Yoshihiro Onouchi

Yoshihiro Onouchi was born in Kyoto, Japan, in 1968. He graduated from the Osaka University School of Medicine in 1994. After a three-year internship at the pediatric department of the Osaka University School of Medicine and its affiliated hospital, he went on to graduate school at the same university and obtained his PhD in 2001. Onouchi came to RIKEN as a research scientist in 2002 after postdoctoral training at the Institute of Medical Science, the University of Tokyo. He was promoted to senior research scientist in 2005. In 2011, he moved to Chiba University as a lecturer and his title at RIKEN changed to visiting researcher. His research focuses on the genetic background of diseases in childhood, especially Kawasaki disease.
