Sep. 19, 2014 Perspectives Biology
Banking on biological tools for research innovation
RIKEN is engaging in international collaborations and targeted research to strengthen repositories of biological materials and information.
The biodiversity of the natural world is an unrivaled source of inspiration and learning. Scientists in the many branches of the life sciences have spent endless hours studying many of the millions of species of microorganisms, plants and animals in order to feed a growing body of knowledge on their cellular, molecular and genetic traits. Through deeper analysis, researchers have found ways to use this knowledge to address emerging societal challenges in health, agriculture and conservation.
Dedicated repositories and distribution centers for biological materials have proven essential to these efforts. By collecting, characterizing and preserving the materials and information that have been captured, identified and developed over decades, resource centers ensure that research in the life sciences remains a progressive endeavor. Without these centers, valuable and irreproducible biological material would be scattered among isolated laboratories and potentially become inaccessible. Resource centers also distribute high-quality materials to laboratories on-demand, a service that saves scientists time and money.
More in one
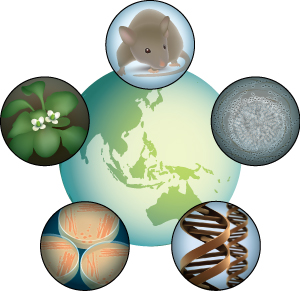 Figure 1: The RIKEN BioResource Center stores five of the most essential categories of biological resources used in research—mice, plants, cells, genes and microbes. © 2014 RIKEN
Figure 1: The RIKEN BioResource Center stores five of the most essential categories of biological resources used in research—mice, plants, cells, genes and microbes. © 2014 RIKEN
The RIKEN BioResource Center (BRC) focuses on five of the most essential categories of biological resources used in research—mice, plants, cells, genes and microbes—and has grown in the past decade to become one of the largest repositories for these resources in the world (Fig. 1). The center preserves more than 4.5 million species, strains, lines and clones, which have been distributed to institutions in over 60 countries and have contributed to more than 400 patent applications. Almost 1,500 papers could not have been published in 2013 without the center’s support.
The BRC is the first repository of its kind in Japan, a responsibility it takes seriously by prioritizing resources developed in the country. Its collection includes microbes, cell lines and blood specimens adopted from other institutes in Japan as well as the Nobel-prize-winning resources of human and mouse induced pluripotent stem (iPS) cells and mouse models developed by Shinya Yamanaka at Kyoto University. And the center has been a core facility of the Japanese National BioResource Project since 2002.
Storing vast quantities of biological material requires a lot of space—a major limiting factor for many centers. Living organisms are particularly expensive to maintain. Therefore, researchers at RIKEN have optimized cryopreservation techniques for freezing sperm and embryos from genetically modified and inbred mouse strains, which can be thawed at a later date to recover live mice with high survival rates and at significant cost efficiencies1. Plant cells are difficult to freeze due to their high water content, but researchers at the BRC have overcome these challenges and introduced an efficient method for cryopreserving plant tissue cultures2.
Knockout genes and universal databases
 Figure 2: Bioresources are preserved in liquid nitrogen tanks at the RIKEN BioResource Center. © 2014 RIKEN
Figure 2: Bioresources are preserved in liquid nitrogen tanks at the RIKEN BioResource Center. © 2014 RIKEN
The coming decade will see dramatic increases in biological resources and the space and computational power required to house and process all the newfound biological organisms and associated information will be huge.
With only a fraction of the world’s species named and classified, there is significant potential for adding new biological ‘products’ to existing collections. Once their genomes have been characterized in full—something that is becoming easier and faster thanks to next-generation gene sequencing technologies—scientists can begin to create an unlimited assortment of transgenic organisms. The further expansion of mutant strains will be facilitated by new gene editing techniques, coupled with gene or allele-specific expression profiling in addition to methods such as the Cre-lox recombination system and optogenetics that can control the timing and expression of target genes.
One destructive technique that is raising particular interest for its speed, ease and precision is CRISPR-Cas. The technique uses clustered regularly interspaced palindromic repeats (CRISPRs), in complex with the protein Cas, to snip a target DNA sequence and even exchange it with another. Unlike traditional gene-targeting techniques that rely on the time-consuming process of homologous recombination in embryonic stem cells, and newer approaches that use proteins to recognize DNA sequences, the CRISPR-Cas system uses single strands of RNA specially designed to ‘knock out’ specific genes. Promisingly, researchers have recently proved the applicability of CRISPR-Cas to gene editing in animal genomes.
Bioresource centers have initiated efforts to keep up with these revolutionizing trends. RIKEN has joined several institutions in a large-scale collaboration known as the International Mouse Phenotyping Consortium (IMPC). The IMPC has set itself the enormous task of generating one mutant mouse for each of the 20,000 estimated genes in the mouse genome. It is now carrying out a pilot study using advanced gene-editing techniques like CRISPR-Cas to create mutant mice at an increased pace than previous methods. By observing how the removal of a single gene affects the structure, development and overall behavior of each mutant mouse, the consortium will be able to create a centralized database of the biological function of each gene, which would be of huge value to scientists. Each mutant mouse sperm and embryo will be preserved in repositories like the BRC.
Besides the IMPC, RIKEN is participating in numerous international efforts that seek to establish a one-stop database for searching and locating biological resources, a process that also involves harmonizing standards for banking, characterization and testing. RIKEN also currently heads the Asian Network of Research Resource Centers, which promotes further coordination in the species-rich region.
Hotspots of innovation
As shores to the foundations of life sciences research, much more will be expected of biological resource centers in the coming years. Rather than acting as static libraries, centers will need to work more closely with scientists by becoming nodes for innovative research. The BRC plans to double the size of its research arm and quicken the pace of research in three key innovation hotspots—iPS cells, the model plant Brachypodium distachyon and bioimaging.
Researchers today can reprogram adult cells into iPS cells and then remodel them into the many specialized cells that constitute our organs. These patient-derived cells are useful tools for studying disease processes and for personalized regenerative medicine. A major collaboration led by the Japanese government is currently underway to develop disease-specific iPS cells, which will be used to investigate pathogenic mechanisms and test new drugs. The project has designated the BRC as its core repository. To add research value, the BRC plans to manipulate the deposited iPS cells to express markers for gene expression such as green fluorescent protein. Given the huge social implications of iPS cell research, ensuring the quality of these cells for future analysis will be critical.
The BRC is also focusing its attention on a new model plant, the grass Brachypodium distachyon, in an effort that parallels RIKEN’s contributions to the extensive infrastructure that now surrounds Arabidopsis thaliana. Brachypodium is a member of the monocot family of flowering plants and is the preferred model for improving our understanding of life-supporting crops such as rice and wheat, as well as species used for biomass production, including switchgrass and zebra grass. In 2013, researchers from the RIKEN Center for Sustainable Resource Science constructed a comprehensive cDNA resource for Brachypodium and subsequently deposited the clones at the BRC3. The BRC has established a platform to utilize the resource and facilitate the dissemination of seeds and genetic alteration technology to the crop research community.
In the field of bioimaging, researchers at RIKEN have developed a chemical mixture for visualizing the whole brain and its cellular circuits, such as neuronal substructures and the activity of fluorescently tagged proteins, with unprecedented detail4. The RIKEN team has also developed a nondestructive method for simultaneously observing DNA methylation, protein localization and RNA expression at specific sites with single-cell resolution, offering new opportunities for studying the epigenetic basis of cellular dynamics during embryonic development, reprogramming and disease5.
By prioritizing key areas of research in this way, and fostering international networks, repositories of biological materials and information such as the BRC are ensuring that life scientists can continue to develop new treatments for diseases and more resilient crops while preserving the world’s most precious biological materials.
References
- 1. Mochida, K., Hasegawa, A., Taguma, K., Yoshiki, A. & Ogura, A. Cryopreservation of mouse embryos by ethylene glycol-based vitrification. Journal of Visualized Experiments 57, 3155 (2011). doi: 10.3791/3155
- 2. Kobayashi, T., Niino, T. & Kobayashi, M. Simple cryopreservation protocol with an encapsulation technique for tobacco BY-2 suspension cell cultures. Plant Biotechnology 22, 105–112 (2005). doi: 10.5511/plantbiotechnology.22.105
- 3. Mochida, K., Uehara-Yamaguchi, Y., Takahashi, F., Yoshida, T., Sakurai, T. & Shinozaki, K. Large-scale collection and analysis of full-length cDNAs from Brachypodium distachyon and integration with Pooideae sequence resources. PLoS ONE 8, e75265 (2013). doi: 10.1371/journal.pone.0075265
- 4. Susaki, E. A., Tainaka, K., Perrin, D., Kishino, F., Tawara, T., Watanabe, T. M., Yokoyama, C., Onoe, H., Eguchi, M., Yamaguchi, S. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014). doi: 10.1016/j.cell.2014.03.042
- 5. Shiura, H., Okamoto, A., Sasaki, H. & Abe, K. Whole-mount MeFISH: A novel technique for simultaneous visualization of specific DNA methylation and protein/RNA expression. PLoS ONE 9, e95750 (2014). doi: 10.1371/journal.pone.0095750
About the Researcher
Yuichi Obata

Yuichi Obata has successfully led the RIKEN BioResource Center for almost a decade. He ensures the strict quality control of biological material stored at the BRC and conducts research in the field of molecular biology and genetics, specifically related to cancer immunology.
