Aug. 13, 2010 Press Release Biology
Scientists clarify structural basis for biosynthesis of mysterious 21st amino acid
Researchers at the RIKEN Systems and Structural Biology Center and the University of Tokyo have clarified the structural basis for the biosynthesis of selenocysteine (Sec), an amino acid whose encoding mechanism offers clues about the origins of the genetic alphabet. The findings deepen our understanding of protein synthesis and lay the groundwork for advances in protein design.
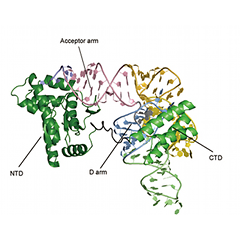 PSTK molecule bound to tRNASec
PSTK molecule bound to tRNASec
One of the most remarkable aspects of translation, the process whereby genetic information is converted into proteins in cells, is its universality: nucleotide triplets ("codons") encode a set of twenty amino acids that form the building blocks for all living organisms. Selenocysteine, the "21st amino acid" whose antioxidant properties help prevent cellular damage, is a rare exception to this rule. Structurally similar to the amino acid serine (Ser) but with an oxygen atom replaced by the micronutrient selenium (Se), selenocysteine is synthesized through a complex juggling of the cell's translational machinery whose mechanisms remain poorly understood.
Central to this multi-step process is a Sec-specific transfer RNA (tRNASec) with an unusual structure that enables it to hijack the "stop codon" UGA to allow incorporation of selenocysteine during protein synthesis. In earlier work, the researchers identified features of tRNASec that differentiate it from other tRNA, notably the peculiar structure of a domain called the D-arm, which appeared to act as an identification marker for recognition by the selenocysteine synthesis machinery. This time, the team analyzed the D-arm's role in the interaction of tRNASec with O-phosphoseryl-tRNA kinase (PSTK), a protein whose selective phosphorylation is essential for selenocysteine encoding.
Using X-ray crystallography, the team showed for the first time that it is the unique structure of the tRNASec D-arm which enables PSTK to distinguish tRNASec from other tRNA. Reported in the August 13th issue of Molecular Cell (online August 12th), the discovery clarifies a pivotal step in selenocysteine biosynthesis, shedding new light on the mysterious 21st amino acid and the elaborate process by which it is created.
Reference
- Shiho Chiba, Yuzuru Itoh, Shun-ichi Sekine and Shigeyuki Yokoyama. Structural Basis for the Major Role of O-Phosphoseryl-tRNA Kinase in the UGA-Specific Encoding of Selenocysteine. Molecular Cell 39: 1-11. August 13, 2010. DOI: 10.1016/j.molcel.2010.07.018
Contact
Shigeyuki Yokoyama
RIKEN Systems and Structural Biology Center (SSBC)
Tel: +81-(0)45-503-9196 / Fax: +81-(0)45-503-9195
Jens Wilkinson
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr@riken.jp
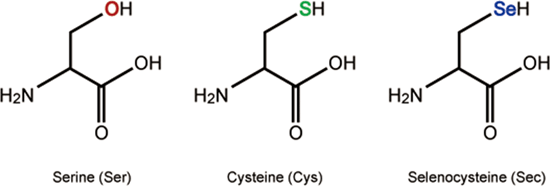
Figure 1: Comparison of selenocysteine (Sec) to the structurally similar amino aids serine (Ser) and cysteine (Cys)
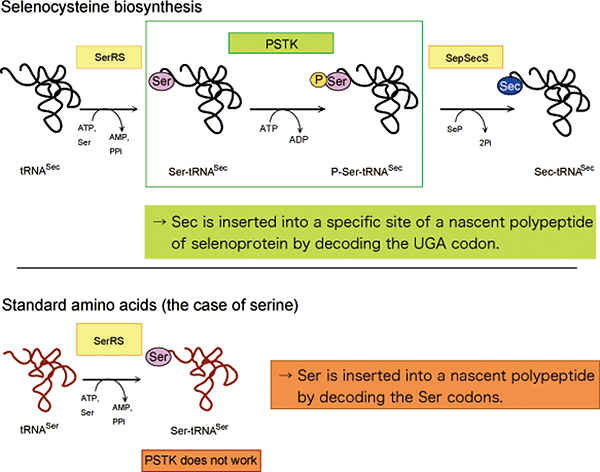
Figure 2: Selenocysteine biosynthesis
Top: tRNASec is first ligated with serine to form Ser-tRNASec. The seryl moiety of Ser-tRNASec is then phosphorylated by PSTK to yield P-Ser-tRNASec, which is converted to Sec-tRNASec and used on the ribosome to insert Sec into a specific site in a nascent polypeptide of selenoproteins.
Bottom: In the case of the standard amino acid serine, tRNASer is ligated with serine and directly used for translation. Ser-tRNASer is not a substrate of PSTK.
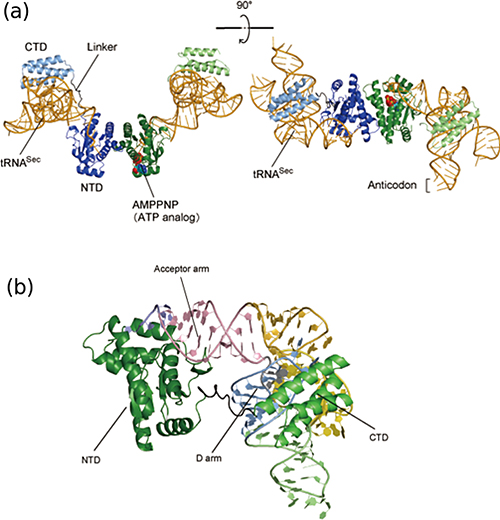
Figure 3: Structure of the tRNASec • PSTK complex
(a) Two PSTK molecules (colored blue and green) interact with each other to form a homodimer. Each PSTK molecule binds a tRNASec molecule. PSTK consists of two independent, linker-connected domains, the N-terminal catalytic domain (NTD) and the C-terminal domain (CTD). These domains independently bind tRNASec.
(b) A close-up view of one of the PSTK molecules bound to tRNASec. The N-terminal domain (NTD) and the C-terminal domain (CTD) of PSTK interact with the acceptor arm (colored pink) and the D arm (light blue), respectively. PSTK does not interact with the tRNASec anticodon complementary to the UGA codon.
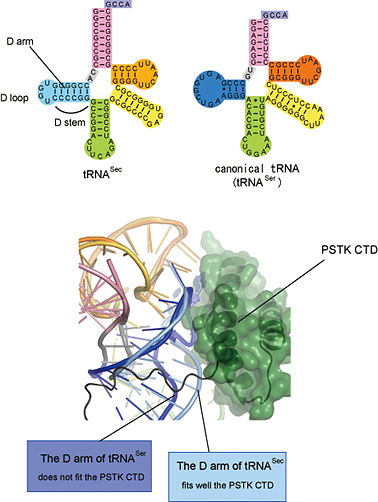
Figure 4: Interaction between the unique D arm of tRNASec and the PSTK CTD
Top: Comparison of the secondary structure of tRNASec to that of a canonical tRNA. The tRNASec D arm consists of a six base-pair stem (D stem) and a four-nucleotide loop (D loop), in contrast with the 3-4 base-pair D stem and the 7-11 nucleotide D loop of the canonical tRNA.
Bottom: The D arm of tRNASec (colored light blue) snugly interacts with the PSTK CTD (green), whereas the D arm of the standard tRNA (blue) does not fit the PSTK CTD.
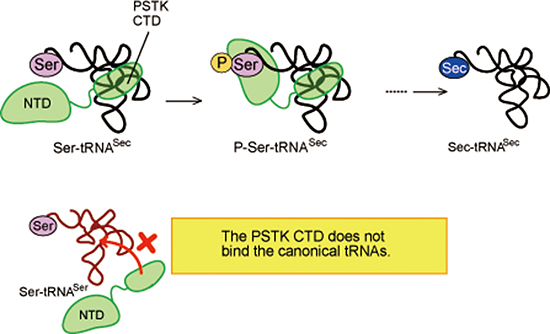
Figure 5: tRNASec recognition by PSTK
The enzymatic activity of PSTK is governed by the specific interaction between its CTD and the unique D arm of tRNASec. The tight binding of the CTD to the D arm ensures that the N-terminal catalytic domain binds to the end of the acceptor arm, where the phosphorylation reaction occurs. In contrast, the CTD does not fit the D arm of canonical tRNAs, and thus PSTK does not act on them, segregating the Sec insertion pathway from the normal amino-acid translation process.
