Jan. 21, 2016 Press Release Biology Medicine / Disease
Genetic variation may explain Asian susceptibility to Kawasaki disease
Scientists from the RIKEN Center for Integrative Medical Sciences (IMS) in Yokohama, Japan, in collaboration with researchers from a number of hospitals around Japan, have found two variations in a gene called ORAI1, one of which may help explain why people of Asian descent are more susceptible to Kawasaki disease, a poorly understand ailment that mostly afflicts young children. The work was published in PLOS ONE.
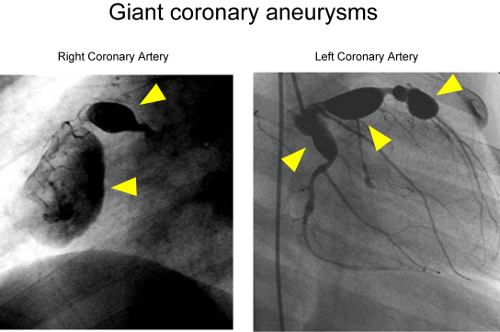
The etiology of Kawasaki disease, which causes inflammation of blood vessels leading to symptoms such as fever and redness of the eyes, lips, and tongue, is a mystery, because it shows seasonal variations—hinting at an infectious or other environmental cause—but is also more prevalent in children of East Asian ancestry—suggesting that there are genetic factors behind it. Kawasaki disease is normally a self-limiting condition, improving over time, but in some cases it is accompanied with potentially fatal aneurysms of the coronary arteries. Previous studies have linked Kawasaki disease to the calcium NFAT pathway, which regulates immune function in response to calcium signals, but which is also known to be involved in the development of the cardiac and other systems. Considering this, the RIKEN-led team decided to focus their work on ORA1, a gene within a chromosomal region where a positive linkage signal has been seen in a previous genome-wide linkage study and had shown to be involved in the activation of the calcium NFAT pathway.
To identify variations that might be associated with susceptibility to the disease, the group examined the genes of 729 Kawasaki disease patients and 1,315 healthy adults. To identify variations that were most strongly associated with the disease, they then looked at variations in a further 1,813 patients and 1,097 non-patients. From these analyses, they identified a single nucleotide polymorphism (SNP), rs3741596, that was much more common in the patients than in the healthy controls. This SNP had not been identified in previous studies on Kawasaki, but intriguingly, it had been found from genomic studies of populations (HapMap) that the variant is common in East Asian populations, and to a lesser extent in African populations, but was very rare elsewhere. They also discovered a second, rare variation that was also highly correlated with the diseases.
According to Yoshihiro Onouchi of the IMS Laboratory for Cardiovascular Disease, who led the study, “It has been suggested that a pathway involved in the immune response is associated with Kawasaki disease, and we knew from our previous work that the calcium NFAT pathway is somehow linked to the disease. It was interesting—and makes a lot of sense—that the common variation we discovered is common in East Asia.”
He continues, “These findings give added credence to the idea that the NFAT pathway is involved in the etiology of Kawasaki disease, and will help us to understand the causes and mechanisms behind it. We hope also that it could lead to treatments for the disease, which in some cases has devastating consequences.”
Reference
- Yoshihiro Onouchi, Ryuji Fukazawa, Kenichiro Yamamura, Hiroyuki Suzuki, Nobuyuki Kakimoto, Tomohiro Suenaga, Takashi Takeuchi, Hiromichi Hamada, Takafumi Honda, Kumi Yasukawa, Masaru Terai, Ryota Ebata, Kouji Higashi, Tsutomu Saji, Yasushi Kemmotsu, Shinichi Takatsuki, Kazunobu Ouchi, Fumio Kishi, Tetsushi Yoshikawa, Toshiro Nagai, Kunihiro Hamamoto, Yoshitake Sato, Akihito Honda, Hironobu Kobayashi, Junichi Sato, Shoichi Shibuta, Masakazu Miyawaki, Ko Oishi, Hironobu Yamaga, Noriyuki Aoyagi, Megumi Yoshiyama, Ritsuko Miyashita, Yuji Murata, Akihiro Fujino, Kouichi Ozaki, Tomisaku Kawasaki, Jun Abe, Mitsuru Seki, Tohru Kobayashi, Hirokazu Arakawa, Shunichi Ogawa, Toshiro Hara, Akira Hata and Toshihiro Tanaka, "Variations in ORAI1 gene associated with Kawasaki disease", PLOS ONE, doi: 10.1371/journal.pone.0145486
Contact
Group Director
Toshihiro Tanaka
Visiting Scientist
Yoshihiro Onouchi
Laboratory for Cardiovascular Diseases
RIKEN Center for Integrative Medical Sciences
Jens Wilkinson
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr@riken.jp