Jul. 6, 2017 Press Release Biology Medicine / Disease
Visualizing whole-body cancer metastasis at the single-cell level
Researchers at the RIKEN Quantitative Biology Center (QBiC) and the University of Tokyo (UTokyo) have developed a method to visualize cancer metastasis in whole organs at the single-cell level. Published in Cell Reports, the study describes a new method that combines the generation of transparent mice with statistical analysis to create 3-D maps of cancer cells throughout the body and organs.
Recent research in optical clearing methods has made it possible to transparentize the bodies and organs of experimental animals. This has led to a new wave of anatomical studies that can combine tissue transparency with sophisticated cell-labeling techniques and light microscopy. Led by Hiroki Ueda at RIKEN QBiC/UTokyo and Kohei Miyazono at the UTokyo, the team has focused their efforts on being able to visualize and profile cancer metastasis throughout the body.
“One of the biggest difficulties in studying cancer,” explains co-senior author Ueda, “is that tumor metastasis is started by just a few metastasized cells. Our new method makes it possible to image the whole body down to the individual cell level, and therefore we can detect cancer at spatial resolutions beyond what is possible using other current imaging techniques.”
The team first focused on finding the best refractive index for their clearing agent, which is called CUBIC. A refractive index is a numerical value that describes how much light is bent as it moves through an object, which can affect the quality of the images that can be obtained from light microscopes. They tested a range of refractive indices and determined which one was the best. Next, when they used the CUBIC method with this refractive index with—termed CUBIC-R—and combined it with different kinds of microscope technology, they were able to detect metastasis of single cells throughout the body and organs of the mouse specimens.
Co-senior author Miyazono notes that this method is a bridge between in vivo imaging of live animals and conventional histology. “Although we cannot apply our new CUBIC-Cancer analysis to live animals, we were able to quantify metastatic cells very early in formation. This will be a very powerful tool for evaluating the effectiveness of anti-cancer drugs.”
The team tested this hypothesis by treating mice with anti-cancer drugs and using CUBIC-Cancer analysis to profile their effectiveness. They were able to detect differences in the total volume and number of cancer colonies throughout the lungs of the mice. The technique was even able to detect individual cancer cells. As co-first authors Kubota and Takahashi explain, “this is very promising because these cells might be dormant or resistant to anti-cancer drugs. As just one surviving cancer cell can lead to tumor metastasis, being able to use CUBIC-Cancer analysis to evaluate drug effectiveness at this level is going to be a very useful and practical application.”
Reference
- Kubota SI, Takahashi K, Nishida J, Morishita Y, Ehata S, Tainaka K, Miyazono K, Ueda HR (2017) Whole-Body Profiling of Cancer Metastasis with Single-Cell Resolution. Cell Reports. doi: 10.1016/j.celrep.2017.06.010
Contact
Group Director
Hiroki Ueda
Laboratory for Synthetic Biology
Cell Design Research Core
RIKEN Quantitative Biology Center
Adam Phillips
RIKEN International Affairs Division
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr@riken.jp
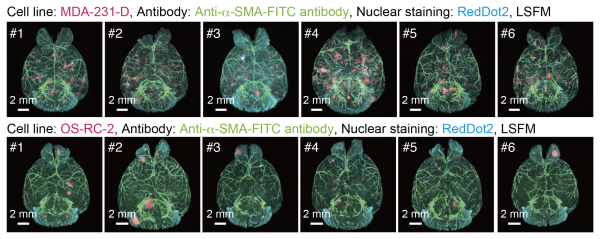
3D analysis of the metastatic patterns in experimental brain metastasis models
Whole-brain imaging of the experimental brain metastasis models with MDA-231-D cells or OS-RC-2 cells in BALB/c-nu/nu mice. The 3D images of the brain samples are shown (upper: MDA-231-D at Day 28, lower: OS-RC-2 at Day 40, cancer cell: mCherry, α-SMA: FITC, nuclei: RedDot2). Animal number of each group is n = 6.
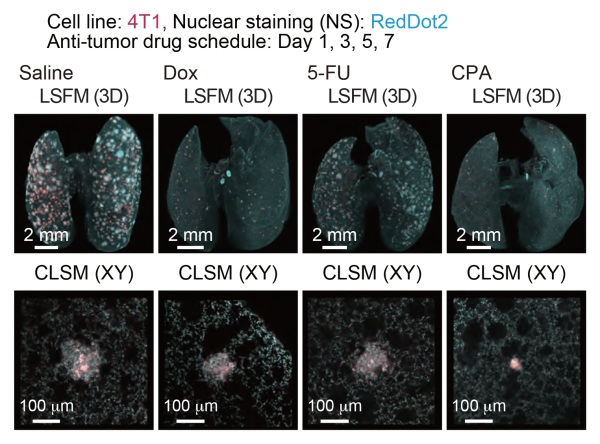
Quantitative evaluation of therapeutic effects of anti-tumor drugs in an experimental lung metastatic model
In vivo therapeutic efficacy of anti-tumor drugs. The 3D images of the lung samples are shown (4T1: mCherry, nuclei: RedDot2).
