Aug. 29, 2018 Press Release Biology
Two genes that regulate how much we dream
Sleep is known to allow animals to re-energize themselves and consolidate memories. Rapid eye movement (REM) sleep, a mysterious stage of sleep in which animals dream, is known to play an important role in maintaining a healthy mental and physical life, but the molecular mechanisms behind this state are barely understood. Now, an international research team led by researchers at the RIKEN Center for Biosystems Dynamics Research (BDR) in Japan has identified a pair of genes that regulate how much REM and non-REM sleep an animal experiences.
Sleep is a universal and vital behavior in animals. In higher vertebrates such as mammals and birds, sleep is classified into two phases, rapid eye movement (REM) sleep and non-REM sleep. During REM sleep, our brain is as active as it is during wakefulness, and this stage is believed to function in memory consolidation. Although our knowledge of the neural mechanisms underlying sleep has gradually advanced, the essential molecular factors that regulate REM sleep are still unknown. Now, however, a research team led by Hiroki Ueda at RIKEN BDR and The University of Tokyo has identified two essential genes involved in the regulation of REM sleep. The amount of REM sleep was drastically decreased down to almost undetectable levels when both genes were knocked out in a mouse model. This study was published by Cell Reportson August 28.
Several past studies have suggested that acetylcholine—the first identified neurotransmitter—and its receptor are important for the regulation of REM sleep. Acetylcholine is abundantly released in some parts of mammalian brain during REM sleep and wakefulness. However, it was unclear which receptor or receptors were directly involved in the regulation of REM sleep due to the complexity of the underlying neural network.
For this study, the researchers used cutting-edge genetic tools to modify mouse genes and conduct genetic screening for factors whose inhibition would cause sleep abnormalities. After knocking out a number of genes encoding various acetylcholine receptors, they found that the loss of two receptors—called Chrm1 and Chrm3—induced a characteristic short-sleep profile. These two receptors are widely distributed in distinct brain regions. The knockout of Chrm1 reduced and fragmented REM sleep, whereas knocking out Chrm3 reduced the length of non-REM sleep. When both genes were knocked out, mice failed almost entirely to experience REM sleep, but survived nonetheless.
“The surprising finding that mice are viable despite the almost complete loss of REM sleep will allow us to rigorously verify whether REM sleep plays a crucial role in fundamental biological functions such as learning and memory” says Yasutaka Niwa, the co-first author of this article.
These findings strongly suggest that these two receptors are essential for sleep regulation, especially REM sleep, and function in different ways. “The discovery that Chrm1 and Chrm3 play a key role in REM sleep opens the way to studying its underlying cellular and molecular mechanisms and will eventually allow us to define the state of REM sleep, which has been paradoxical and mysterious since its original report,” Ueda says.
Additional data is available for readers who have an extensive interest in the phenotype of the mice who lost all their REM sleep.
Reference
- Yasutaka Niwa*, Genki N. Kanda*, Rikuhiro G. Yamada*, Shoi Shi, Genshiro A. Sunagawa, Maki Ukai-Tadenuma, Hiroshi Fujishima, Naomi Matsumoto, Koh-hei Masumoto, Mamoru Nagano, Takeya Kasukawa, James Galloway, Dimitri Perrin, Yasufumi Shigeyoshi, Hideki Ukai, Hiroshi Kiyonari, Kenta Sumiyama, Hiroki R. Ueda# (*Co-first,#Lead contact), "Muscarinic acetylcholine receptors Chrm1 and Chrm3 are essential for REM sleep", Cell Reports, 10.1016/j.celrep.2018.07.082
Contact
Team Leader
Hiroki Ueda
Laboratory for Synthetic Biology
RIKEN Center for Biosystems Dynamics Research
Masataka Sasabe
RIKEN International Affairs Division
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr[at]riken.jp
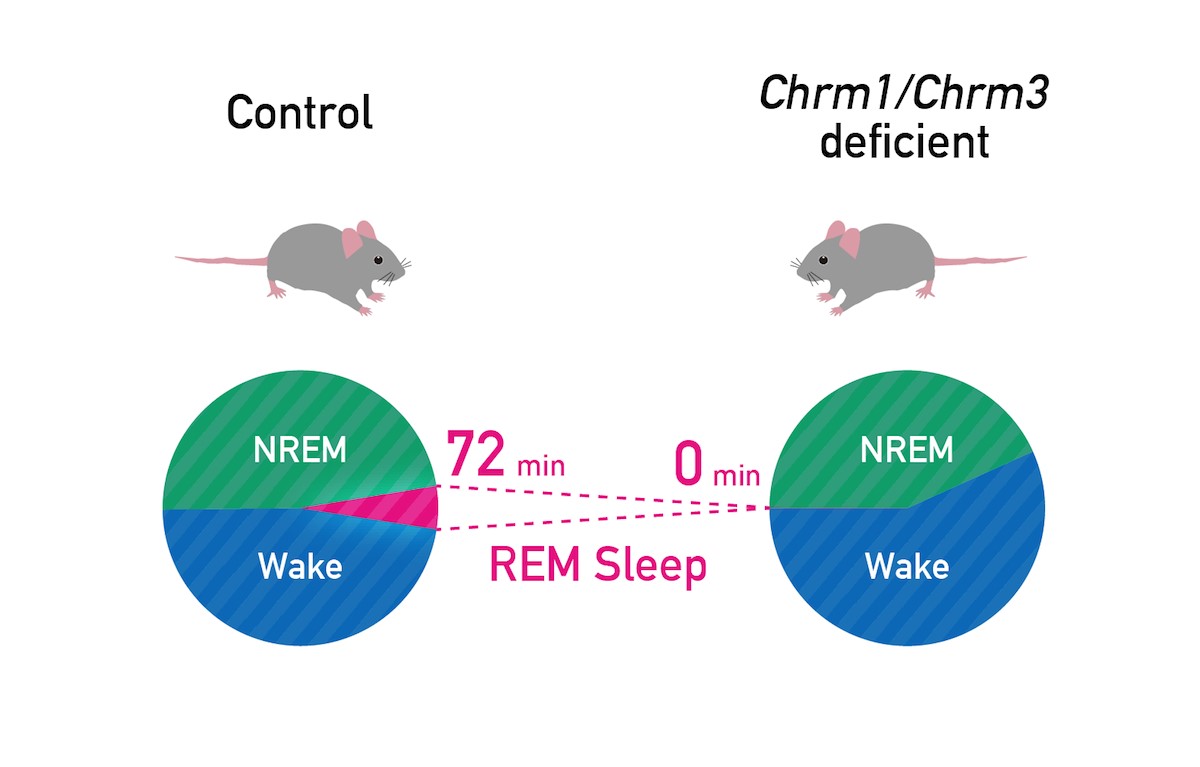
Double KO of Chrm1 and Chrm3
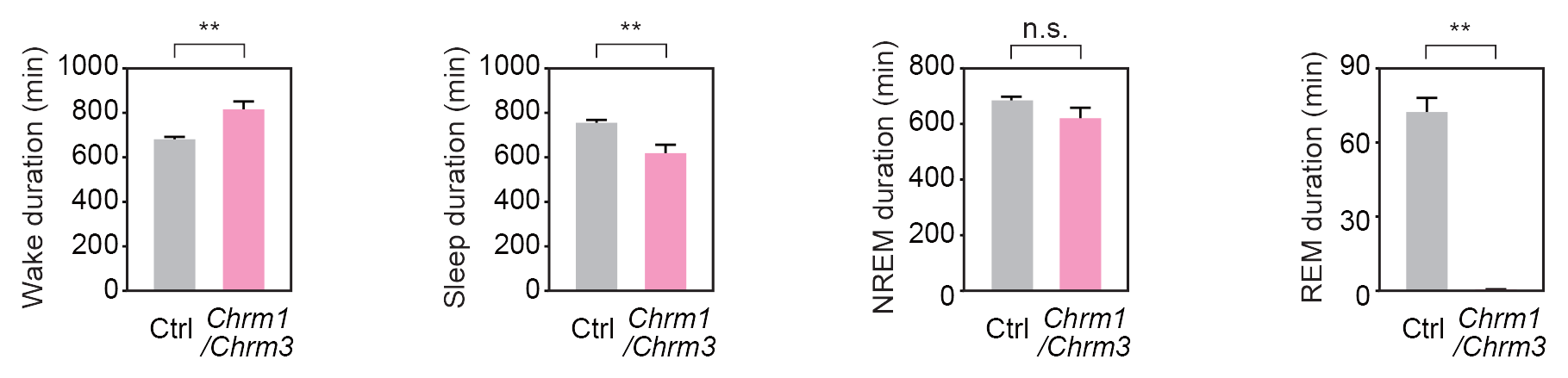
Sleep phenotypes of ES-cell-derived Chrm1/3 double KO mice

Receptors Chrm1 and Chrm3 regulate REM sleep
Acetylcholine receptors Chrm1 and Chrm3 (illustrated as Sandman gnomes 1 and 3) are essential for REM (dream) sleep. In this visual description of the study, the receptors help a mouse dream about a sleeping cat and Michel Jouvet (1925-2017), a pioneer of REM sleep studies. © Hiroko Uchida.
