Aug. 14, 2020 Research Highlight Biology
Probe detects the destruction of defective mitochondria
A fluorescent probe that works in both live and fixed cells can provide key insights into neurodegenerative and other disorders
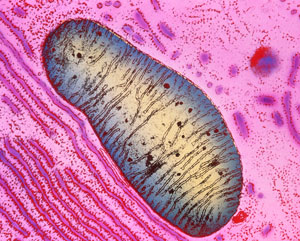 Figure 1: Colored transmission electron micrograph of a single mitochondrion in a human pancreas cell. RIKEN researchers have developed a fluorescent probe that can detect the programmed death of defective mitochondria in the lysosomes.© K.R. PORTER/ SCIENCE PHOTO LIBRARY
Figure 1: Colored transmission electron micrograph of a single mitochondrion in a human pancreas cell. RIKEN researchers have developed a fluorescent probe that can detect the programmed death of defective mitochondria in the lysosomes.© K.R. PORTER/ SCIENCE PHOTO LIBRARY
A versatile probe that can detect with pinpoint accuracy the programmed destruction of defective mitochondria—the powerhouses of cells—has been developed by RIKEN researchers1. They used it to show that damaged mitochondria in dopamine-producing neurons fail to be destroyed in mice with a condition resembling Parkinson’s disease.
Mitochondria are organelles that generate most of the chemical energy our cells need to function. But when cells are stressed, mitochondria can malfunction and produce highly reactive oxygen radicals, which damage cells. Thus, cells routinely weed out and destroy defective mitochondria by assigning them to lysosomes, which function as the waste-disposal system of cells, breaking down unwanted components.
If this selective elimination of dysfunctional mitochondria—known as mitophagy—fails, it can lead to various diseases. There is thus much interest in monitoring mitophagy in cells.
Fluorescent probes have been developed that can detect mitophagy. But some can only be used in living cells, while others are vulnerable to destructive processes that do not involve lysosomes.
Now, Atsushi Miyawaki of the RIKEN Center for Brain Science and co-workers have developed a new fluorescent probe that can be used in both living and fixed cells and is highlighted specifically in lysosomes.
Their probe contains two parts: one that can withstand the enzymes in the lysosome and another that is destroyed by them. Thus, by monitoring the color of the probe’s fluorescence, the researchers could detect when a mitochondrion had entered a lysosome. Unlike their previous mitophagy probe, the new probe is sensitive to the degrading enzymes in lysosomes and acidity, so it works even in fixed cells where lysosomes are no longer acidic.
 Hiroyuki Katayama (left), Atsushi Miyawaki and their co-workers have developed a fluorescent probe that can accurately detect mitophagy in live and fixed cells and used it to show that dopamine-producing neurons in a mouse model of Parkinson’s disease failed to eliminate defective mitochondria.© 2020 RIKEN
Hiroyuki Katayama (left), Atsushi Miyawaki and their co-workers have developed a fluorescent probe that can accurately detect mitophagy in live and fixed cells and used it to show that dopamine-producing neurons in a mouse model of Parkinson’s disease failed to eliminate defective mitochondria.© 2020 RIKEN
The team used the probe to investigate Parkinson’s disease—a neurodegenerative disease that causes shaking, muscle stiffness and progressive difficulties with movement.
Using a mouse model of Parkinson’s disease, the researchers found that neurons that produced the neurotransmitter dopamine failed to eliminate defective mitochondria, but other neurons that did not produce dopamine did. Since Parkinson’s disease is characterized by a dopamine deficiency in the brain, this suggests that the inability of dopamine-producing neurons to perform mitophagy could be a major factor in the disease.
By collaborating with researchers from the pharmaceutical company Takeda, Miyawaki’s team identified a compound that can induce the destruction of damaged mitochondria. Such compounds could help to treat Parkinson’s disease in the future.
The probe is promising for advancing research into other diseases. “Since many other neurodegenerative disorders involve mitophagy, our probe can contribute to their study,” says Miyawaki. “Furthermore, diseases in other organs involve oxidative stress and hence mitophagy. We’re currently using our probe to look at heart disease.”
Related contents
References
- 1. Katayama, H., Hama, H., Nagasawa, K., Kurokawa, H., Sugiyama, M., Ando, R., Funata, M., Yoshida, N., Homma, M., Nishimura, T. et al. Visualizing and modulating mitophagy for therapeutic studies of neurodegeneration. Cell 181, 1176–1187.e16 (2020). doi: 10.1016/j.cell.2020.04.025
