Jul. 6, 2007 Research Highlight Biology Chemistry
Working shifts could aid disease detection
Specific, fluorescent dyes could light the way in recognizing genetic disorders resulting from small DNA sequence variations
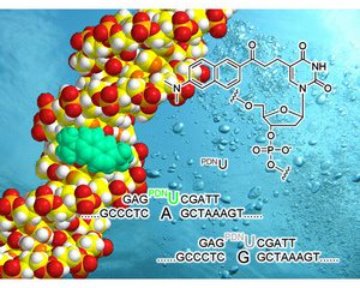 Figure 1: A graphical representation of the fluorescent DNA probe in action.
Figure 1: A graphical representation of the fluorescent DNA probe in action.
A new method to detect small changes in human genes could lead the way in personalized medicine. The most common type of variation in our genes is a single difference in one of the nucleotide building blocks of the DNA sequence, known as single nucleotide polymorphism (SNP—pronounced ‘snip’). Scientists believe differences in SNPs reveal an individual’s susceptibility to disease, meaning accurate analysis of SNPs would play a key role in diagnostics. These small SNP variations can account for as little as 0.1 per cent of a genome sequence.
SNP diagnostics have recently attracted much attention and several strategies to identify SNPs have been developed in the past few years. Existing methods are often limited by the need to identify large DNA sequences. Now, a team of Japanese researchers led by Akimitsu Okamoto from the RIKEN Frontier Research System, Wako, has used derivatives of the fluorescent dye PRODAN to correctly identify SNPs quickly and efficiently1.
PRODAN, a well-known fluorophore, absorbs and emits light at different wavelengths depending on the polarity of its environment. Okamoto reasoned that similar dyes, which include differing nucleotide components, could be incorporated into DNA structures and ‘report back’ differences in the microenvironment. Such changes in the microenvironment would likely be the result of small changes in the DNA structure and allow detection of sequence variations (Fig. 1). The team synthesized four variants of the dye, so that all combinations of base matches and mismatches could be investigated.
Once incorporated in a DNA sequence under a variety of conditions, the team calculated the differences between the wavelengths absorbed and emitted by the dye. These differences are known as Stokes shifts. The researchers detected a small Stokes shift when DNA base pairs matched correctly, but a larger shift when there was a mismatch. Therefore, by using various PRODAN-labeled DNA dyes, single nucleotide alterations could be detected. “The use of this DNA probe makes it possible to judge the type of base located at a specific site on the target DNA, simply by mixing the DNA and the dye together. This method is a very powerful assay that does not require enzymes or time-consuming steps, and avoids errors,” says Okamoto.
Okamoto believes that this method of detection is very promising and is working towards making the system suitable for every day use. “I think SNP chips using our probe would make important contributions to cancer diagnosis,” he enthuses.
References
- 1. Tainaka K., Tanaka K., Ikeda S., Nishiza K., Unzai T., Fujiwara Y., Saito I. & Okamoto A. PRODAN-conjugated DNA: Synthesis and photochemical properties. Journal of the American Chemical Society 129, 4776–4784 (2007). doi: 10.1021/ja069156a
