Jun. 6, 2014 Research Highlight Biology
Genetic ‘junk’ may have its uses
Genome filler could have an important role in maintaining stem cell potency
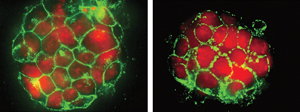 Figure 1: Fluorescence microscopy images of mouse embryonic stem cells (left) and induced pluripotent stem cells (right) showing the expression of a key marker of stem cell pluripotency (green). © 2014 Alexandre Fort, RIKEN Center for Life Science Technologies
Figure 1: Fluorescence microscopy images of mouse embryonic stem cells (left) and induced pluripotent stem cells (right) showing the expression of a key marker of stem cell pluripotency (green). © 2014 Alexandre Fort, RIKEN Center for Life Science Technologies
A fundamental process in biology is the transcription of genes into various types of RNA. Many RNAs code proteins that are essential for cell function. However, the body of transcripts, or transcriptome, of mammalian cells also contains a diversity of noncoding RNAs, many of which are derived from self-replicating DNA sequences known as retrotransposons that ‘jump’ around the genome. Generally considered to be genetic filler, the biological functions of retrotransposons and their transcripts, particularly in mammalian stem cells, remains largely unknown.
Piero Carninci, Alexandre Fort, Kosuke Hashimoto and colleagues from the RIKEN Center for Life Science Technologies and RIKEN Center for Integrative Medical Sciences have now led research that has revealed that a number of retrotransposon-derived transcripts are implicated in maintenance of the undifferentiated or pluripotent state of stem cells1.
Retrotransposons can quickly make copies of themselves in the genome and are responsible for evolutionary increases in the size of the genome, as well as the introduction of genetic variations. Among the many types of retrotransposons, long terminal repeat (LTR) retrotransposons share common features with retroviruses, leading many scientists to regard them as inactive remnants of ancient viral invasions that have been incorporated into host genomes during evolution.
To better understand the noncoding part of the stem cell transcriptome, Carninci’s team performed deep profiling of nuclear and cytoplasmic transcriptomes from a representative set of human and mouse stem cell lines, including embryonic and induced pluripotent stem cells, using four complementary high-throughput sequencing technologies (Fig. 1). The research team identified thousands of non-annotated transcripts—genomic sequences not annotated as transcribed (or RNA-producing) regions. By characterizing these transcripts and highlighting those found only in stem cells, the researchers revealed that an important fraction of these non-annotated stem transcripts (NASTs) originated in LTR retrotransposons.
Manipulation of four LTR-associated NAST candidates induced the differentiation of pluripotent iPS cells. The findings suggest that LTR retrotransposons and their transcripts are likely to be involved in the maintenance of pluripotency.
“Our work has just begun to unravel the scale of unexpected functions carried out by retrotransposons and their derived transcripts in stem cell biology,” says Carninci. “We were extremely surprised to learn from our data that what were once considered genetic ‘junk’ are in reality symbiotic elements that work closely with other genes to maintain embryonic and induced pluripotent stem cells in their undifferentiated state. This is quite different from the image given by textbooks.”
References
- 1. Fort, A., Hashimoto, K., Yamada, D., Salimullah, M., Keya, C. A., Saxena, A., Bonetti, A., Voineagu, I., Bertin, N., Kratz, A. et al. Deep transcriptome profiling of mammalian stem cells supports a regulatory role for retrotransposons in pluripotency maintenance. Nature Genetics 46, 558–566. doi: 10.1038/ng.2965
