Understanding how complex biological molecules evolved has long been an important part of the quest to learn how life developed on earth. Some complex proteins are made of structurally similar units in symmetrical arrangements, and it is commonly believed that they evolved by the duplication and fusion of genetic material encoding these units.
One class of complex proteins is the beta-propeller family, which is composed of many similar but different domains arranged in a ring around a central point, with an overall propeller-like shape. In nature, we know of beta-propeller proteins with between four and ten blades.
It was commonly believed, however, after some failed attempts to make such proteins, that a perfectly repeating protein would be unable to fold. This raised questions about whether multi-domain proteins could evolve from simple duplication and fusion events, since the intermediate forms would be made up of perfectly repeating units.
This puzzle is exactly what Arnout Voet, a Foreign Postdoctoral Researcher at the Structural Bioinformatics Team in the RIKEN Center for Life Science Technologies, decided to take on. Imagine, he wondered, if we could perform the supposedly impossible task of creating a symmetrical protein, made up of blades like a propeller arranged in a regular order?
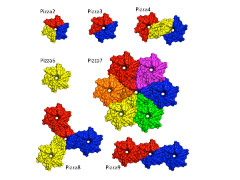
Voet, under the supervision of Kam Zhang of CLST and Jeremy Tame of Yokohama City University, began by using a novel computer-aided protein design procedure in which they "reverse engineered" the evolutionary process to create a perfectly symmetrical protein. The sequence that led to a structure with the lowest calculated energy was then turned into an artificial gene, and the symmetrical six-bladed protein was made and purified. The team studied this protein using a number of techniques to show that the result of the computation was in perfect accord with the experimental results. Because of the shape, they named this protein Pizza6.
They showed that a propeller protein with six identical blades can indeed fold, and be highly stable. Their designer proteins show that beta-propeller proteins could well have evolved through gene duplication and fusion.
Though the work was motivated by the quest to understand how life evolved, it could also have important significance for bionanotechnology. Through the work with computational modeling, the group has shown a new way to create highly stable, symmetrical protein building blocks which can self-assemble into higher-order structures. These results could lead to the rational design of multi-functional biomaterials, and the method could be used with other templates and symmetries as desired.
Voet says, "Artificial protein design remains a new field, yet it promises all sorts of new biomaterials if we can master it. The work we did was not only fun, but covered different aspects of protein science from evolution, sequence analysis and computer modelling to a half-dozen techniques in biophysics. We made something that did not exist before, but which may one day be useful."
