Apr. 29, 2014 Press Release Biology Medicine / Disease
Viral "parasites" may play a key role in the maintenance of cell pluripotency
In a study published in Nature Genetics, scientists from the RIKEN Center for Life Science Technologies in Japan, in collaboration with the RIKEN Center for Integrative Medical Sciences, the University of Copenhagen and the Joint Genome Institute (Walnut Creek, California) have discovered that "jumping DNA" known as retrotransposons—viral elements incorporated into the human genome—may play a key role in the maintenance of pluripotency, the ability of stem cells to differentiate into many different types of body cells.
This story is part of a fundamental rethinking taking place in genomic science. In 2009, members of the FANTOM Consortium project reported that an important fraction of mammalian transcriptomes—meaning the RNA transcribed from the genome—consists of transcripts derived from retrotransposon elements, vestiges of ancient retroviruses from the same family as HIV that have in the past been considered to only parasite the genome. However, the biological function of these "jumping DNA"-associated RNA transcripts remained unknown.
In the current study on embryonic stem (ES) cells and induced pluripotent stem (iPS) cells using four high-throughput methods including cap analysis gene expression (CAGE), the researchers found that thousands of transcripts in stem cells that have not yet been annotated are transcribed from retrotransposons, presumably to elicit nuclear functions. These transcripts were found to be expressed in stem cells, but not differentiated cells. Importantly, the work showed that several of these transcripts are involved in the maintenance of pluripotency, since degrading several of them using RNA interference caused iPS cells to lose their pluripotency and differentiate.
These transcripts appear to have been recruited, surprisingly both in the human and mouse genome, where they are used to maintain the pluripotency of stem cells. Somehow, organisms including humans appear to have recruited viral elements into their genome in a way that helps to maintain the pluripotency of stem cells that allow them to regenerate. Why this is so remains a mystery for future investigation.
Although the results of the study cannot be put directly into application in regenerative medicine, knowing that retrotransposon elements are essential in the transcriptional control of iPS and ES cells is an essential clue for solving the puzzle of how to create better types of cells in future regenerative medicine studies.
"Our work has just begun to unravel the scale of unexpected functions carried out by retrotransposons and their derived transcripts in stem cell biology. We were extremely surprised to learn from our data that what was once considered genetic 'junk', namely ancient retroviruses that were thought to just parasite the genome, are in reality symbiotic elements that work closely with other genes to maintain iPS and ES cells in their undifferentiated state. This is quite different from the image given by textbooks that these genomic elements are junk," explains Dr. Piero Carninci, senior investigator of the study.
The study was funded by a JSPS (Japan Society for Promotion of Science) grant for Next Generation World-Leading Researchers.
Reference
- Alexandre Fort, Kosuke Hashimoto, Daisuke Yamada, Md Salimullah, Chaman A. Keya, Alka Saxena, Alessandro Bonetti, Irina Voineagu, Nicolas Bertin, Anton Kratz, Yukihiko Noro, Chee-Hong Wong, Michiel de Hoon, Robin Andersson, Albin Sandelin, Harukazu Suzuki, Chia-Lin Wei, Haruhiko Koseki, The FANTOM5 Consortium, Yuki Hasegawa, Alistair R.R. Forrest & Piero Carninci. "Deep transcriptome profiling of mammalian stem cells supports a regulatory role for retrotransposons in pluripotency maintenance". Nature Genetics, 2014, doi:10.1038/ng.2965
Contact
Alexandre Fort or Piero Carninci
Division of Genomic Technologies
RIKEN Center for Life Science Technologies
Jens Wilkinson
RIKEN Global Relations and Research Coordination Office
Tel: +81-(0)48-462-1225 / Fax: +81-(0)48-463-3687
Email: pr[at]riken.jp
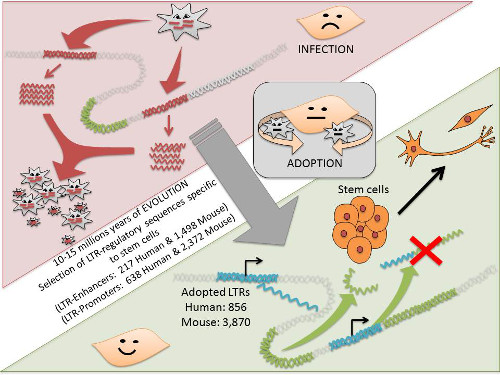
Evolution of retrotransposon domestication in the genome
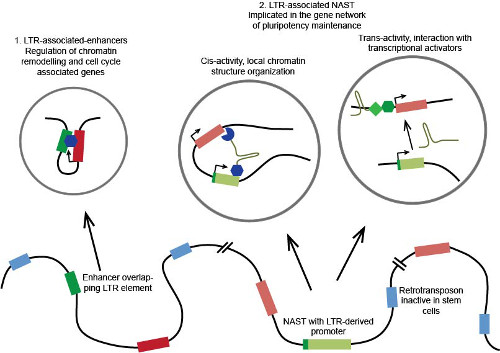
Putative molecular functional mechanism of domesticated LTR regulatory elements
