Nov. 30, 2018 Research Highlight Biology
Movement patterns of receptors can indicate their activation status
Monitoring the movement behavior of an essential protein family could greatly speed up drug discovery
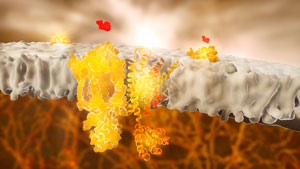 Figure 1: Adrenergic receptors (or adrenoceptors) are a class of G protein-coupled receptors (GPCRs). A RIKEN-led team has found that monitoring the diffusion of such receptors can be used to determine whether or not they are activated by a compound. © RAMON ANDRADE 3DCIENCIA/SCIENCE PHOTO LIBRARY
Figure 1: Adrenergic receptors (or adrenoceptors) are a class of G protein-coupled receptors (GPCRs). A RIKEN-led team has found that monitoring the diffusion of such receptors can be used to determine whether or not they are activated by a compound. © RAMON ANDRADE 3DCIENCIA/SCIENCE PHOTO LIBRARY
The discovery of new drugs for conditions ranging from depression to diabetes to cancer could be accelerated thanks to a promising strategy identified by RIKEN researchers for quantifying the activation of a key class of proteins1.
Many leading drugs for a wide variety of conditions act on a single class of proteins, known as G protein-coupled receptors (GPCRs). The human genome encodes roughly 800 GPCRs, which coordinate a wide array of essential cellular functions.
To design a new drug for a GPCR, it is first necessary to establish whether a particular compound activates that receptor. This is a daunting task, because each GPCR triggers a distinct chain of signaling events, which can be difficult to dissect.
Yasushi Sako, who heads the Cellular Informatics Laboratory at RIKEN, and his co-workers thus set out to identify other indicators that might reveal whether a given receptor is switched on or off.
Masataka Yanagawa, a scientist in Sako’s team, employed a sensitive imaging technique that allowed him to track the behavior of individual fluorescently labeled GPCR molecules within the cell membrane. Over the course of his observations, he was struck by an apparent link between the activation state of individual receptor molecules and the rate at which they diffused within the fluid membrane environment.
 Masataka Yanagawa (left), Yasushi Sako (right) and their co-workers have obtained new insights into the physical behavior of an essential protein family that could greatly accelerate the drug discovery process. © 2018 RIKEN
Masataka Yanagawa (left), Yasushi Sako (right) and their co-workers have obtained new insights into the physical behavior of an essential protein family that could greatly accelerate the drug discovery process. © 2018 RIKEN
“If we can find a general relationship between the diffusion and function of GPCRs, we could estimate a drug’s effect just by monitoring the diffusion behavior of the receptor molecule,” Yanagawa notes.
The researchers focused on a GPCR known as metabotropic glutamate receptor 3 (mGluR3), which contributes to communication between neurons and has been linked to disorders such as schizophrenia. Their imaging data revealed striking changes in this receptor’s diffusion behavior when it was switched on versus when it was turned off—activating signals produced by stimulatory ‘agonist’ molecules generally slowed the receptor down or even immobilized it, whereas chemical treatments that inactivated mGluR3 generally produced faster diffusion within the membrane.
Remarkably, a similar pattern held for a wide variety of other GPCRs. “Despite having considerably different amino acid sequences, all the GPCRs we tested showed a significant decrease in their mobility on stimulation,” says Yanagawa. This suggested a pattern of behavior that may be common across all such proteins.
Sako’s group, in collaboration with Masahiro Ueda’s group at the RIKEN Center for Biosystems Dynamics Research, has recently developed a high-speed imaging platform2, which could help researchers to exploit this response to quickly identify promising drug candidates that turn clinically relevant GPCRs on or off.
Related contents
- Turning off cellular signaling offers new drug hopes
- Lighting up drug discovery
- Imaging lipid rafts reveals some surprises
References
- 1. Yanagawa, M., Hiroshima, M., Togashi, Y., Abe, M., Yamashita, T., Shichida, Y., Murata, M., Ueda, M. & Sako, Y.Single-molecule diffusion-based estimation of ligand effects on G protein–coupled receptors. Science Signaling 11, eaao1917 (2018). doi: 10.1126/scisignal.aao1917
- 2. Yasui, M., Hiroshima, M., Kozuka, J., Sako, Y. & Ueda, M. Automated single-molecule imaging in living cells. Nature Communications 9, 3061 (2018). doi: 10.1038/s41467-018-05524-7
