Nov. 7, 2023 Research Highlight Biology
Uncovering the cell’s read–write mechanism for gene expression instructions
Researchers have uncovered how instructions for gene expression are relayed
The ‘read–write’ mechanism by which cells replicate and use chemical instructions for expressing genes has been uncovered by RIKEN researchers1.
The quality and quantity of gene expression correlates not only with instructions by transcription factors but also with chemical modifications to the various histone proteins, which provide a scaffold for DNA in the chromosomes.
Scientists have long argued whether these modifications to histones are the epigenetic cause for activating gene expression. And, if that is the case, how they activate gene expression and are maintained during the process of mitosis, in which a cell divides into two daughter cells.
“Whether histone modifications are the epigenetic cause for gene expression has remained a hypothesis because no one had ever seen whether histone modifications self-replicate,” explains Takashi Umehara of the RIKEN Center for Biosystems Dynamics Research.
To explore this question, Umehara and his team focused on a protein known as p300/CBP—an enzyme that can both introduce and bind to acetyl-group modifications (acetylations) on histone proteins. Specifically, the researchers were interested in specific acetylations on the histone H3–H4 complex to which p300/CBP binds. These acetylations are known to activate gene expression in nearby DNA sequences.
But H3–H4 is just one component of a larger ‘nucleosome’ assembly, which also includes the histone H2B–H2A complex. All of these various histones can carry distinct acetylation patterns, and the causal relationships between their acetylations have not been well understood.
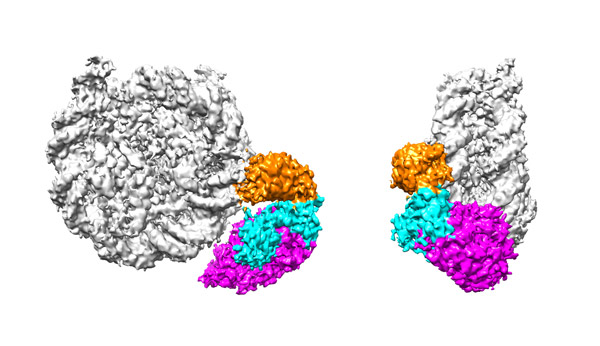
Figure 1: Two views of how p300/CBP (colored) propagates acetylation of histones in the nucleosome complex (gray). The orange ‘read’ domain recognizes and binds to acetylations on H3–H4, while the magenta ‘write’ domain transcribes acetylations from H3–H4 to H2B–H2A. © 2023 RIKEN Center for Biosystems Dynamics Research
Now, Umehara and colleagues have developed an experimental technology that allowed them to generate histones with acetylations at defined sites. They then monitored how p300/CBP interacts with and acetylates a nucleosome containing these selectively acetylated human histones.
The team found that p300/CBP recognizes and binds to specific acetylation marks on the H3–H4 complex. The enzyme then replicates acetylation marks to unacetylated sites of H3–H4, while also transcribing them from H3–H4 to H2B–H2A within the same nucleosome. Since this newly acetylated H2B–H2A complex is more likely to be stripped from the nucleosome, a model emerges in which it finally instructs which genes to be transcribed by the cellular transcription machinery.
These results provide an unprecedented glimpse into how p300/CBP inherits acetylation marks to newly divided cells and utilizes those marks epigenetically for gene expression. “I could never have imagined such an elegant yin–yang mechanism for the inheritance and expression of epigenetic information,” says Umehara.
Umehara’s team now aims to explore how well conserved these processes are across non-animal species, including yeast and plants.

A team led by Takashi Umehara (second row; first on the left) and Masaki Kikuchi (first row, second from the right) has uncovered how instructions for gene expression get relayed to the goal. © 2023 RIKEN
Related contents
- Mapping the chromatin landscape reveals determinants of placental stem cell identity
- Inheriting acquired traits requires trailblazer modifications to unfertilized eggs
- Gene-reading enzyme razes and rebuilds DNA-winding structures in its path
Rate this article
Reference
- 1. Kikuchi, M., Morita, S., Wakamori, M., Sato, S., Uchikubo-Kamo, T., Suzuki, T., Dohmae, N., Shirouzu, M. & Umehara, T. Epigenetic mechanisms to propagate histone acetylation by p300/CBP. Nature Communications 14, 4103 (2023). doi: 10.1038/s41467-023-39735-4
