Research related to COVID-19 (Updated April 1, 2022)
Overview
In 2020 the novel coronavirus (COVID-19) began to spread worldwide. RIKEN launched a special research project in April to help tackle this global crisis. Since then, in collaboration with other research institutes, universities and companies, RIKEN has been conducting vigorous research and development and making its facilities, including the supercomputer Fugaku, available to the public in order to respond quickly and flexibly to various needs. Through this project, we have succeeded in developing more efficient and rapid virus detection methods and therapeutic drugs, and in generating data for the development of effective infection-prevention measures.
RIKEN will continue to contribute to overcoming COVID-19 by maximizing the use of its resources and sharing new knowledge with the world to sustain human life and society.
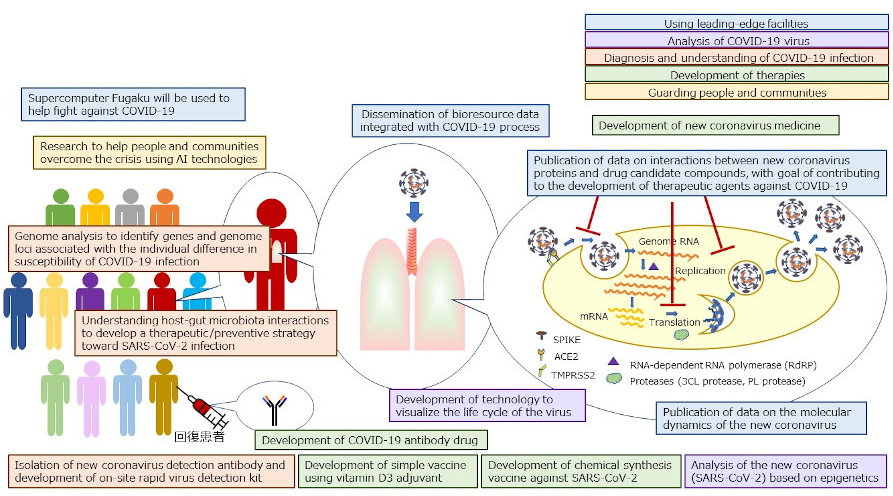
Research related to COVID-19
Related movies on YouTube
- RIKEN's response to COVID-19 (COVID-19 research, #1)
- A new type of vaccine (COVID-19 research, #2)
- How does COVID-19 spread in air?
- New COVID-19 workplaces
Research Themes
- Publication of data and research using leading-edge large-scale joint-use facilities
- Development of diagnostic methods
- Research for the development of therapies
- Research to help people and communities overcome the crisis
- Basic and other research
Publication of data and research using leading-edge large-scale joint-use facilities
SPring-8/SACLA call for research proposals related to COVID-19
SPring-8/SACLA has opened a call for research proposals related to COVID-19 to contribute to efforts to combat the disease.
Publication of data on interactions between new coronavirus proteins and drug candidate compounds, with goal of contributing to the development of therapeutic agents against COVID-19
Scientists have used the fragment molecular orbital method (FMO) to calculate molecular interactions between the proteins of the new SARS-CoV-2 virus and candidate drug compounds, and published this data on the FMO database (FMODB) on April 17 to allow researchers around the world to freely use the data for drug discovery.
Dissemination of bioresource data related to COVID-19 and other infectious diseases
We are working to develop an integrated database of relationships between bioresources and biological processes of viral infection and aggravation of infectious diseases including COVID-19, to enable prompt data dissemination on newly developed bioresources useful for infectious disease research.
Development of diagnostic methods
Development of methods to rapidly diagnose COVID-19 at high sensitivity without amplification
Based on a "digital detection method of nucleic acids" developed by RIKEN, work is being carried out to rapidly develop a system to detect novel coronavirus-derived RNA rapidly and at high sensitivity, without the need for amplification.
Predicting COVID-19 symptoms and outcomes before infection, in a precise and personalized way
Susceptibility and resistance to SARS-CoV-2 depends on multiple factors, including the central role played by innate and acquired immunity, which are in turn influenced by previous vaccinations, such as BCG, and infections from other coronaviruses. The gut microbiota also helps shape the immune system, and it can be damaged by the inappropriate use of antibiotics. Why some people rapidly progress to respiratory failure in COVID-19 is poorly understood, with multiple associations observed between outcomes and hypertension, diabetes, obesity, and cardiovascular disease. Several genes have been identified from research on other coronaviruses, including ACE2 and TMPRSS2, which are directly involved in viral infection. Through international collaboration, we will unravel these phenotypic and genetic associations to provide a better understanding of specifically why and how COVID-19 leads to mortality in some patients.
Development of data-driven infection-control strategies based on stratification of preclinical populations
COVID-19 is characterized by a diversity of symptoms, with the majority of infected people remaining asymptomatic or mild, while in some cases the disease becomes severe or fatal. In comparison to the substantial data collected in hospitals after the onset of disease, data from before the onset has not been sufficiently accumulated, and no methodology has been established to assess the risk of developing illness and severe illness for each individual in advance. In this study, we will collect saliva and nasopharyngeal swab samples from thousands of healthy and asymptomatic individuals by exploiting the large-scale social PCR testing system that we have already started to build. We will comprehensively measure human- and microbial-derived DNA and RNA contained in these samples, and develop a method to evaluate the individual risk of developing illness and severe illness before the onset of disease through an approach that combines machine learning with statistical and mathematical modeling. We will also develop a model for predicting epidemic trends that takes into account the diversity of disease risk among individuals, with the aim of creating personalized infection-control strategies that determine when, who, and what interventions are most effective.
Virus detection device using metamaterial color structure
Using the resonant interaction between nanoscale metal structures and light waves, it is possible to create artificial color pigments. By applying this technique to immunochromatography, we will develop an ultra-high-sensitive virus detection device that can detect antigen or protein molecules from a virus as their color changes.
A pooled assay that enables detection and identification of new virus variants
We will develop a pooled approach that enables detection and identification of new SARS-CoV-2 variants leveraging our microfluidic technologies.
Rapid and sensitive on-site measurement of antibodies against the COVID-19 virus
We are working to develop a microarray system for detecting viruses that can rapidly and with high sensitivity measure the amount of antibodies (antibody titer) against the novel coronavirus (SARS-CoV-2) in human blood. With this system, we aim to make it possible to efficiently and precisely test the effectiveness of SARS-CoV-2 vaccinations on site, a task which will become increasingly important at medical facilities.
More information is available on the press release page for this project.
Research for the development of therapies
Development of therapeutic drugs targeting Main protease / Papain-like protease
When viruses invade cells they produce a long protein (polyprotein) based on the genetic information in their own RNA. The polyprotein molecules are cleaved into more than a dozen fragments by the virus-derived “main protease” and “papain-like protease,” which are then used for viral proliferation and for the formation of new virus particles. By targeting this mechanism, we aim to develop drugs that inhibit the function of those proteases and suppress viral replication in cells.
Development of therapeutic drugs targeting TMPRSS2
The surface of the COVID-19 virus has protrusions called “spike proteins” that bind to the ACE2 receptor on the surface of human cells, forming a precursor. The binding site of the precursor is modified by TMPRSS2, an enzyme expressed in human cells, allowing the virus to invade. Drugs that inhibit the action of TMPRSS2 could effectively block the entry of viruses into cells. Since TMPRSS2 is a human enzyme, it is expected that inhibitors will not be affected by viral mutations.
Immunotherapy using artificial adjuvant vector cells (aAVC)
aAVCs are cells loaded with glycolipids, which activate innate immunity, and antigens, which activate acquired immunity. Until now, aAVCs have been developed for cancer immunotherapy. A RIKEN team hypothesized that using aAVCs might make it possible to suppress COVID-19 with a double-pronged attack from both innate and acquired immunity. This could be done by administering aAVCs carrying spike proteins and envelope proteins on the virus surface as antigens to patients.
Drug discovery based on structural analysis of biomolecules related to COVID-19
This project aims to clarify the quaternary structures of various proteins involved in SARS-CoV-2 infection, taking them as drug targets for COVID-19. Some of the target proteins have quaternary structures consisting of multiple proteins such as non-structural proteins (NSPs) or have large degrees of freedom. Depending on their properties, the project is using cryo-electron microscopy and nuclear magnetic resonance (NMR) to obtain structural information useful for design.
Drug discovery research based on molecular simulations of biomolecules related to COVID-19 and artificial intelligence (AI)
Many of the candidate proteins for COVID-19 drug targets have flexible protein surfaces and the dispersive force interactions are important. Preliminary studies have shown that conventional docking methods do not have sufficient ability to predict activity. This project will work to select candidate compounds by applying a new activity prediction method with higher accuracy combining FMO (quantum chemical calculation) and AI.
Development of COVID-19 antibody drug
Toward the development of antibody treatment against COVID-19, we are aiming to isolate a human monoclonal neutralizing antibody against SARS-CoV-2 and its variants.
Protein dynamics using rigidity-based analysis in the coronavirus
We are conducting rigidity-based analysis on the conformational dynamics of proteins in SARS-CoV-2 for understanding COVID-19 corona virus molecular mechanisms to aid in therapeutic design.
Development of chemical synthesis vaccine against SARS-CoV-2
Our team has developed a new technology "mMAP" (mutation compatible multiple antigen peptides) which could help to overcome the problems of virus mutation and antibody dependent enhancement. A COVID-19 vaccine using this technology is currently under development.
Development of simple vaccine using vitamin D3 adjuvant
Our goal is to establish a safe and simple vaccine for COVID-19. Herd immunity requires the global community to be immunized simultaneously. We will work to develop a needle-less vaccine system by taking an advantage of our new adjuvant discovered at RIKEN.
Screening of drug candidate compounds for COVID-19 in large databases using scalable similarity searches
We are screening drug candidate compounds for COVID-19 from a large database using scalable similarity searches.
Structural study of innate immune protein respond to viral RNA for drug development against SARS CoV-2
To understand the activation mechanism of innate immune proteins that recognize foreign RNAs, we will perform structural analysis of RNA-protein complexes by cryo-electron microscopy (CRYO ARM 300).
Establishment of animal models for development of new drugs and vaccines against COVID-19
For development of new drugs and vaccines, we are establishing animal models that will show severe symptoms of the coronavirus infection as infected humans.
Development of an all-round antibody to block all variants the novel coronavirus
Based on the fact that variant coronaviruses can have stronger infectivity and weaker reactivity toward neutralizing antibodies, we will establish therapeutic antibodies which attack not the virus itself but the molecule on host cells critical to viral infection and invasion, and which are able to block infection by any novel coronavirus variant.
Research to help people and communities overcome the crisis
Development of remote communication and dialogue support system for healthcare during standby at home due to COVID-19 epidemic
Staying at home due to the COVID-19 epidemic may raise the risk of frailty and cognitive decline in older adults. Once older adults with frailty and/or cognitive decline are infected, it is difficult for them to return from hospitals home or to elderly care facilities, leading to pressure on medical facilities. We are developing a remote communication and dialogue support system for maintaining the cognitive and physical health of older adults.
Analyze of hate speech and disinformation related to COVID-19
We are collecting and analyzing instances of abusive languages such as hate speech and disinformation related to the COVID-19 pandemic using knowledge and techniques of language philosophy and natural language processing.
ELSI and further perspectives of on-line initial medical examination
We are classifying and reevaluating the ELSI (Ethics, Legal and Social Issues) of on-line initial medical examinations.
Investigation of influences caused by teleworking on workers and possible improvements
Teleworking conditions have caused various problems on the environment of workers, e.g., physical fatigue and work postures, mental stress from lack of conversations, hard situation with online meeting in a small room through small screen, ambiguity between "ON" and "OFF" work periods, and how to make "OFF" situations clear. We are investigating elements and components of these issues and will propose possible ways to improve the situation.
Machine learning based prediction of severity of COVID-19 infection using health insurance claims data and information from medical checkups
We are developing concise and rapid machine learning based methods for the prediction of disease severity especially among younger COVID-19 infected persons using nationwide large health insurance claims data and medical checkup data.
Countermeasures to COVID-19 by decentralized data sharing, analysis and utilization integrating AI/ICT/HPC
To help prevent the spread of COVID-19 infections and address social and economic impacts, we are developing coordination among AI / ICT / HPC using a secure personal-data management technology (PLR: Personal Life Repository) for visualizing infection risk, recommending low-risk routes, simulating infection transmission and economic activities, planning countermeasures, and so forth.
Machine learning based individual treatment optimization of online notice delivery for behavioral change
It is well known in preventive medicine and behavioral economics that traditional information delivery is not effective for behavioral change. To aid effective information delivery for infection prevention of COVID-19 such as keeping away from high-risk behaviors, we are developing methods for machine learning-based individual treatment optimization of online notice delivery based on various studies in preventive medicine and behavioral economics and carrying out experiments based on the methods developed.
Technical, social and legal measures to prevent the spread of COVID-19 infections
COVID-19 infectious diseases require comprehensive measures to prevent the spread of the disease, including, laws, social conditions and technical measures not to mention medical measures such as drugs and vaccines. Legal restrictions, self-restraint, social pressure, affordance, nudge, etc. can be considered as promising legal, social means and privacy protection, but these are not effective on their own, and how they can be combined is pursued in a form of Japanese-English joint research activities.
Study of Social Connection and Loneliness in Post-COVID-19 Society
Self-isolation and social distancing amid the COVID-19 pandemic has had a lasting influence on the mental and physical well-being of people around the world through the effects of loneliness or lack of adequate social connections and interactions. To deal with these social issues of wellbeing, we are examining the quantity as well as quality of social connections, which are indispensable for sustaining healthy lives. In order to elucidate the causation of loneliness affecting mental and physical wellbeing, we are conducting international surveys to elucidate the social structure surrounding the state of loneliness. By doing this, we will not only reveal the circumstances of loneliness within one country but the universal pattern of social structures that cause a state of loneliness. The research will also help identify which individuals in the society are suffering most severely and urgently need help. The results of this study will aid policy decision-making in local and national governments.
A Study of Remote Psychotherapy for Family Support under the COVID-19 Crisis
The COVID-19 pandemic has made it difficult to provide face-to-face psychotherapy to support family relationships. By verifying the effectiveness of remote psychotherapy in parent-child relationships, we are aiming to develop a technology and specific tips about remote therapy that can be used even after the pandemic.
Basic and other research
Development of technology to visualize the life cycle of the virus
RIKEN is developing technology to visualize the virions in infected cells and bio-specimens using fluorescent proteins and single-chain anti-virus antibodies.
Genome analysis of SARS-CoV-2
Tracing of virus genome evolution is important for controlling COVID-19. We have initiated a comprehensive virus genome study of SARS-CoV-2 in Japan virus using RIKEN's genome research infrastructure.
Development of an academic search support system related to COVID-19
In order to support biomedical research on COVID-19 with information science, we are developing an AI system that automatically analyzes abstracts of academic papers related to COVID-19 and facilitates semantic-based multiple paper search mechanisms.
Analysis of the new coronavirus (SARS-CoV-2) based on epigenetics
We are analyzing the effects of methyltransferases that cause RNA methylation such as N6-methyladenosine of SARS-CoV-2 RNA on virus replication and host response by machine learning, and exploring the possibilities to use it as a target for COVID-19 therapy.
Genome analysis to identify genes and genome loci associated with the individual difference in susceptibility to COVID-19 infection
The individual differences in susceptibility to COVID-19 infection can be partially explained by individual difference in genome sequences. We will collect samples in collaboration with various hospitals to analyze the genomic DNA of patients. We will also work with an international consortium.
Understanding host-gut microbiota interactions in order to develop a therapeutic/preventive strategy toward SARS-CoV2 infection
(1) In collaboration with the University of Tokyo and Keio University, we will collect fecal samples from patients who recovered from COVID-19 and search for bacterial strains that can enhance antibody response against the virus. (2) Since we recently identified serine protease-degrading bacterial strains, we will evaluate the effect of these strains against intestinal coronavirus infection.
Study of more severe COVID-19 in the post-vaccine era
To study COVID-19 severity caused by virus escape from vaccines, we will collect and analyze comprehensive data on newly developing mutant strains and patients infected with mutant strains, and validate our findings using animal models. In addition, we will mimic the severity of COVID-19 using humanized mice to advance the search for therapeutic agents.
Data assimilation research with COVID-19 infectious disease models
A standard infectious disease model known as the SIR model will be extended to consider the unique features of COVID-19. Data assimilation will be applied to the extended SIR model, providing useful information for decision-making such as restrictions and vaccination plans.
Mechanism of Inactivation of New Coronaviruses by Ultraviolet Irradiation
The inactivation of the SARS-CoV-2 virus using ultraviolet light, which can be applied to diverse spaces, object surfaces, and liquids, has attracted much attention, and the effectiveness of some wavelengths of UV light has been reported. However, the mechanism through which UV light inactivates SARS-CoV-2 has not been elucidated. In this study, we aim to elucidate the infectivity of the virus and the mechanism of its inactivation using ultraviolet light with a wavelength of 253.7 nm, which is known to have antiviral effects and is the most inexpensive, most easily obtained and practically used wavelength.
Link to paperDevelopment of laser virus inactivation technology
A light-based partition against the novel coronavirus that inactivates coronavirus-containing droplets that pass through a sheet by creating a sheet-like beam with an ultraviolet lamp in the UV-C region or a UV-laser system.
Identification of a cross-reactive epitope for memory killer T cells that react to the novel coronavirus
Researchers have identified a cross-reactive epitope in the spike protein of SARS-CoV-2 that can induce specific killer T cells in individuals with the HLA-A24 type. They showed that the killer T cells also respond to a portion of seasonal coronaviruses that cause the common cold. The epitope would be useful for diagnosis and the development of new vaccines for COVID-19.
